Abstract
Over half of hepatocellular carcinoma (HCC) cases diagnosed worldwide are in China1,2,3. However, whole-genome analysis of hepatitis B virus (HBV)-associated HCC in Chinese individuals is limited4,5,6,7,8, with current analyses of HCC mainly from non-HBV-enriched populations9,10. Here we initiated the Chinese Liver Cancer Atlas (CLCA) project and performed deep whole-genome sequencing (average depth, 120×) of 494 HCC tumours. We identified 6 coding and 28 non-coding previously undescribed driver candidates. Five previously undescribed mutational signatures were found, including aristolochic-acid-associated indel and doublet base signatures, and a single-base-substitution signature that we termed SBS_H8. Pentanucleotide context analysis and experimental validation confirmed that SBS_H8 was distinct to the aristolochic-acid-associated SBS22. Notably, HBV integrations could take the form of extrachromosomal circular DNA, resulting in elevated copy numbers and gene expression. Our high-depth data also enabled us to characterize subclonal clustered alterations, including chromothripsis, chromoplexy and kataegis, suggesting that these catastrophic events could also occur in late stages of hepatocarcinogenesis. Pathway analysis of all classes of alterations further linked non-coding mutations to dysregulation of liver metabolism. Finally, we performed in vitro and in vivo assays to show that fibrinogen alpha chain (FGA), determined as both a candidate coding and non-coding driver, regulates HCC progression and metastasis. Our CLCA study depicts a detailed genomic landscape and evolutionary history of HCC in Chinese individuals, providing important clinical implications.
Similar content being viewed by others
Main
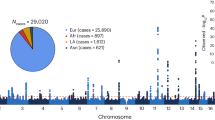
Previous genomic analyses of HCC in Chinese individuals are limited in cohort size and focus mainly on the exome11,12,13,14, precluding detailed investigations at the whole-genome level. Recently, the Pan-Cancer Analysis of Whole Genomes (PCAWG) Consortium analysed the genomic complexity of cancer at a considerable scale4,5,6,7,8. Nevertheless, the relatively shallow sequencing depth could not fully resolve the subclonal structure of the HCC genome. Here, in the CLCA, we performed deep whole-genome sequencing (WGS) analysis of 494 HCC tumours (average depth, 120×), as well as of the matched control blood samples (average depth, 36×). Our cohort comprised 427 men (86.4%) and 67 women (13.6%). In comparison to the PCAWG-HCC (n = 248) cohort, the CLCA cohort had higher proportions of HBV infection (94.5% versus 30.6%) and Edmondson–Steiner grades 3 and 4 (85.6% versus 12.1%), but lower proportions of hepatitis C virus (HCV) infection (2.6% versus 55.6%), alcohol drinking (26.7% versus 58.1%) and smoking (36.8% versus 53.6%) (Extended Data Fig. 1a,b, Supplementary Table 1 and Supplementary Note 1). These statistics represent the epidemiology of the Chinese population with liver cancer, highlighting the necessity of the current study. After stringent quality control, a total of 9,287,828 somatic mutations was identified, with a median of 13,735.5 mutations and 95 nonsynonymous mutations for each tumour (Fig. 1). We also performed RNA sequencing (RNA-seq) analysis of 239 tumours from this cohort (Supplementary Table 2).
a, The research strategy. The diagram was created using BioRender. WT, wild type. b, The candidate driver landscape of the CLCA. The top two graphs show the number of all mutations and nonsynonymous mutations identified in each tumour, followed by annotation of clinical variables. BCLC, Barcelona Clinic Liver Cancer staging system. In total, 23 candidate drivers identified in coding regions and 31 candidate drivers identified in non-coding regions are listed, and the mutational frequency (%) is shown next to the gene IDs. The mutation types are indicated on the right and n denotes the number of drivers in the category. lncP, lncRNA promoter; NBNC, double negative for HBV and HCV; NA, not available. Orange gene symbols indicate previously undescribed drivers identified in the CLCA. Underlined drivers are those identified as a driver in different forms. Group 1 had drivers in both coding and non-coding regions, whereas group 2 had drivers only in non-coding regions. Tumours in group 3 had no identified drivers but other somatic mutations. The number of individual tumours included is denoted for groups 1–3. The bar plot on the left shows the clonal and subclonal mutational frequencies of each gene. Statistical analysis was performed using two-sided Fisher’s exact tests with the Benjamini–Hochberg multiple-hypothesis test. Q values are shown next to the bars. A threshold of Q < 0.1 was used for significance.
Candidate coding and non-coding drivers
We identified 23 candidate coding drivers, including TP53, CTNNB1 and ALB (Fig. 1 and Supplementary Note 2). CTNNB1 mutations were mutually exclusive to either TP53 or AXIN1 mutations (Extended Data Fig. 1c), consistent with HCC in European individuals15. Compared with other cohorts5, six previously undescribed candidate coding drivers were identified in the CLCA, including FGA, HNF1A, PRDM11, CDKN1B, BMP5 and ECHS1 (Extended Data Fig. 1d–g). The mutational frequency of TP53 is significantly higher in the CLCA compared with either PCAWG-HCC or TCGA-HCC. By contrast, the mutational frequencies of six previously undescribed candidate coding drivers were comparable across the three cohorts, indicating the prevalence of these candidate drivers.
A total of 31 candidate non-coding drivers was identified, including six promoters, eight long non-coding RNAs (lncRNAs), four lncRNA promoters, five 5′ untranslated regions (UTRs) and eight 3′ UTRs. Five genes were determined as driver events in different forms, indicating convergent evolution. FGA (encoding fibrinogen alpha chain) was determined as both a candidate coding and non-coding (3′ UTR) driver. Mutations in the 3′ UTR of FGA were not enriched for 2–5 bp indels (Q = 0.73) and are therefore not related to the transcription-associated indel signature16. With the exception of all three non-coding drivers reported by PCAWG-HCC, including the TERT promoter, lncRNA NEAT1 and lncRNA promoter of RMRP, all other 28 events (90.3%) were previously undescribed candidate non-coding drivers. These results confer a rich resource to investigate the contributions of non-coding mutations during hepatocarcinogenesis.
Clonality of candidate drivers
Ten candidate drivers showed significant clonality enrichment of mutations, including two coding and eight non-coding drivers (Fig. 1). Two coding drivers, TP53 and ALB, were enriched with clonal mutations. By contrast, 62.5% (5 out of 8) of non-coding drivers were enriched with subclonal mutations, including the promoters of ZNF595, KCNJ12 and OR2A7, and lncRNA and lncRNA promoter of Z95704.4. No significant association between tumour purity and the percentage of clonal drivers was observed across our cohort (Extended Data Fig. 1h), showing that our clonality analysis is not confounded by tumour purity. The identification of subclonal non-coding drivers highlighted the strength of high-depth WGS data in investigating the non-coding genome, partially explained the low number of non-coding drivers identified in previous low-depth WGS studies, and motivated us to systematically investigate the subclonal events in our cohort. Furthermore, a ratio value of mutated nonsynonymous (dN) and synonymous (dS) sites (dN/dS) of higher than 1 for all mutations was observed for both clonal and subclonal coding drivers (Extended Data Fig. 1i), confirming that these drivers are shaped by positive selection, consistent with previous pan-cancer analyses17,18,19.
SBS_H8 is a novel signature
We identified 17 single-base substitution (SBS), 3 doublet-base substitution (DBS) and 8 small insertion-and-deletion (ID) signatures (Extended Data Figs. 1j and 2–4). In comparison to COSMICv3.2, five signatures were novel (Supplementary Table 3 and Supplementary Note 3) containing one SBS signature: SBS_H8; two DBS signatures, DBS_H1 and DBS_H2; and two ID signatures, ID_H3 and ID_H8. DBS_H1 consisted mainly of [C/G/T]C>NN mutations. This signature was found in most tumours and correlated with age as well as other age-related signatures (Extended Data Fig. 3d,f). ID_H8 showed mostly 1 bp cytosine deletions and thymine insertions. It was exclusively found in SBS_H3-positive (COSMIC SBS24) tumours and correlated with SBS24 (Extended Data Fig. 3e,g), suggesting its relevance to aflatoxin exposure.
Notably, SBS_H8 was dominated by T>[A/C] mutations with significant transcriptional strand bias (Fig. 2a–c). Although the pattern of T>A mutations in SBS_H8 was similar to that of aristolochic acid (AA)-related COSMIC SBS22, SBS_H8 also contained a substantial proportion of T>C mutations (21.3%), together leading to an overall cosine similarity of 0.71 between SBS_H8 and SBS22. The low pentanucleotide context cosine similarity of 0.61 further supported that SBS_H8 was a novel signature rather than a combination of SBS22 and other signatures (Extended Data Fig. 3b). SBS_H8 was present in 57.1% (282 out of 494) of CLCA cases, suggesting the prevalence of this previously undescribed signature of HCC in Chinese individuals. High co-occurrence between SBS_H8 and SBS_H2 (SBS22) indicated that the aetiological factor of SBS_H8 might often co-exist with AA. SBS_H8 is present in only 1 out of 326 (0.31%) PCAWG-HCC cases and potentially in chronic liver disease20. These results supported the existence of this signature and its enrichment in HCCs in Chinese individuals.
a–c, Comparison of the mutational profile (a), transcriptional strand bias (b) and pentanucleotide context of T>A mutations (c) of SBS_H8, SBS_H2 and AA-exposed cell lines. Cosine similarity to COSMIC SBS22 is denoted. Statistical analysis was performed using two-sided binomial tests with Benjamini–Hochberg correction for multiple comparisons. Trans, transcribed strand; Untrans, untranscribed strand. d,e, The mutational profiles of the signatures DBS_H2 (d) and ID_H3 (e), both related to AA. f, The correlation between the numbers of mutations associated with SBS_H2, DBS_H2 and ID_H3. The grey plane is the linear regression plane with projection lines showing residuals (red, positive; blue, negative).
As for AA, we not only found the well-established SBS_H2, but also identified two previously undescribed types of AA signatures—DBS_H2 and ID_H3 (Fig. 2d,e). DBS_H2 consisted primarily of TA>NT, TC>AA, TG>AN and TT>AA mutations. ID_H3 showed mainly 1 bp and 2 bp deletions in short repeats. Both DBS_H2 and ID_H3 were almost exclusively found in SBS_H2-positive (SBS22) tumours and were highly correlated with SBS_H2 activity (Fig. 2f). To test whether SBS_H2, DBS_H2 and ID_H3 are directly caused by AA exposure, we treated two cancer cell lines, MCF-10A and HepG2, with sublethal concentrations of AA1 (the major component of AA). The mutational spectrum of each clone showed the presence of SBS_H2, DBS_H2 and ID_H3 (Supplementary Fig. 1), confirming that these mutational signatures can be caused by AA exposure. These findings complemented the AA signature spectrum and revealed the diverse paths of AA mutagenesis. However, notably, SBS_H8 was not found in the mutational spectrum of AA1-treated cell clones (Fig. 2a–c), which further supported that SBS_H8 was not associated with AA exposure.
Unsupervised hierarchical clustering based on mutational signatures classified 494 tumours into 5 clusters (Extended Data Fig. 3h and Supplementary Note 4). SBS_H8 contributed most to cluster V, which was enriched with CTNNB1 mutations (Extended Data Fig. 3i,j). Higher percentages of SBS_H8 were significantly associated with poorer prognosis (Extended Data Figs. 3k and 5a), implying that the underlying aetiology of SBS_H8 might be a carcinogen of the liver. We also analysed the contribution of mutational processes to driver genes and hotspot mutations (Extended Data Fig. 4). Focusing on SBS_H8, JAK1 and CTNNB1 were the top coding drivers and the ALB promoter was the top non-coding driver. Multiple mutation hotspots of CTNNB1, JAK1S729C and TP53H193R were affected by SBS_H8. Moreover, multiple hotspots of TP53 were associated with aflatoxin, while the TP53H179L hotspot was associated with AA exposure. SBS_H8, as well as other signatures related to exogenous factors such as SBS_H2 (AA), SBS_H3 (aflatoxin), DBS_H2 (AA), ID_H3 (AA), SBS_H10 (tobacco) and ID_H8 (aflatoxin), were enriched for clonal mutations compared with subclonal mutations, suggesting that they occurred at earlier stages of tumorigenesis.
HBV integration in ecDNA
Our deep WGS data enabled a comprehensive profiling of genomic rearrangements, including copy-number alterations (CNAs), structural variations (SVs), HBV integrations, extrachromosomal circular DNA (ecDNA) and three forms of clustered alterations—kataegis, chromothripsis and chromoplexy (Extended Data Figs. 5 and 6 and Supplementary Note 5). ecDNA was detected in 27.3% of CLCA tumours (Fig. 3a and Supplementary Table 4), significantly higher than that reported in PCAWG-HCC (13.1%, P = 3 × 10−4; two-sided Fisher’s exact test)21. A total of 76 oncogenes was detected in ecDNA, including HCC driver genes such as MYC (Fig. 3b and Extended Data Fig. 5d). Oncogenes in ecDNA had higher copy numbers and elevated gene expression compared with their counterparts not in ecDNA (Extended Data Fig. 5e,f). The presence of ecDNA was associated with a poor prognosis (Fig. 3c and Extended Data Fig. 5a). Notably, we identified ecDNAs incorporating HBV segments (HBV-ecDNA) in seven patients (Fig. 3d–f) affecting well-known oncogenes such as TERT. HBV segments in ecDNA showed an elevated number of copies, as well as increased expression levels. Despite the fact that HBV-TERT integration has been identified in HCC, our results demonstrated that these integrations can exploit the circular structure of ecDNA and therefore amplify to hundreds of copies. The existence of ecDNA was successfully validated (Fig. 3g). Collectively, these results suggest that ecDNA-based amplification22 may have an important role in HBV-associated HCC.
a, The proportion of different amplicons across the CLCA cohort. Circular, breakage–fusion–bridge (BFB), heavily rearranged and linear, and no focal somatic copy-number amplification detected (fSCNA) amplicon categories are shown. b, The top frequently amplified genes detected in ecDNA. c, Progression-free survival (PFS) of patients in the CLCA stratified by the existence of ecDNA. Statistical analysis was performed using log-rank tests. d,e, Comparison of the copy number (d) and RNA expression (e) of HBV between circular amplicons and other amplicons. For the box plots, the centre line shows median, the box limits indicate the upper and lower quartiles, and the whiskers extend to 1.5× the interquartile range; data beyond the end of the whiskers are outlying points that are plotted individually. n denotes biologically independent samples. Statistical analysis was performed using two-sided Student’s t-tests. TPM, transcripts per million. f, Two representative ecDNA amplicons involving HBV segments detected in two patients. g, CIRCLE-seq reads supporting the structure of ecDNA. Chr., chromosome.
Subclonal catastrophic events
Clustered mutational processes, including chromothripsis23, chromoplexy7,24 and kataegis25, are genomic alterations that are often generated in a single catastrophic event. These alterations are often described as clonal events and support the punctuated evolution of tumours24,25. Whether these clustered alterations could be subclonal events and occur late during tumour evolution remains less explored. We investigated the clonal status of these events with our high-depth WGS data of the CLCA (Extended Data Fig. 6).
We observed chromothripsis in 30.2% of cases (Supplementary Table 4), comparable to that of PCAWG-HCC (32.2%)26. Among those, 61% of high-confidence events affected multiple chromosomes (for example, CLCA_0119), whereas 22% affected only a single chromosome (for example, CLCA_0090) (Fig. 4a). Chromoplexy was observed in 10.1% of CLCA cases; 8.3% of cases contained a single event (such as CLCA_0489) and 1.8% contained multiple events (for example, CLCA_0232) (Fig. 4b). In total, 364 kataegis events were identified in 33.6% of CLCA cases, and 14.6% of cases had multiple kataegis events. We observed the occurrence of kataegis and oscillations in copy-number states, suggesting that localized hypermutation could be associated with regional SVs and chromothripsis27 (Fig. 4c and Extended Data Fig. 7a). Kataegis events were highly enriched in cases with APOBEC signatures (Extended Data Fig. 5g). In total, 46 (13%) kataegis events occurring in 32 cases (6.5%) were subclonal events (Fig. 4d). This result was distinct to that reported by PCAWG-HCC, in which all kataegis events were clonal events, suggesting that kataegis may be subclonal and occur late during hepatocarcinogenesis. In silico analysis further showed that the detected number of kataegis events increased along with the sequencing depth (Fig. 4e), corroborating that our high-depth WGS enabled the detection of subclonal kataegis events. Furthermore, timing analysis showed that 15.1% of kataegis, 67.2% of chromothripsis and 62.7% of chromoplexy events were determined to be subclonal events, respectively (Fig. 4f). Although all of these forms of clustered alterations tended to be clonal rather than subclonal, the broad distribution of odds ratios suggests that these events could occur at various timings during tumorigenesis (Fig. 4g).
a, Circos plots for chromothripsis events. CN, copy number. b, Circos plots for chromoplexy events. Arcs in the same colour denote regions that are involved in the same chromoplexy event. c, Rainfall plots for kataegis events and related SVs and CNAs. BAF, B allele frequency. d, The clonal status composition of kataegis events. Mixed events are indicated in grey. e, The clonal status of kataegis events. Top, the cancer cell fraction distribution of non-kataegis and kataegis mutations. Bottom, the detected kataegis events at different sequencing depths (simulated in silico). f, The timing of three types of clustered alteration events. g, The relative odds of clustered alterations being clonal or subclonal are shown with bootstrapped 95% confidence intervals (top). Bottom, the relative odds of the events being early or late clonal are shown as above.
Pervasive non-coding drivers
Reconstruction of the evolutionary history of the CLCA categorized 44.98% point mutations as subclonal, in contrast to that of 8% in PCAWG-HCC (Extended Data Fig. 7b, Supplementary Table 5 and Supplementary Note 6). In comparison to candidate coding drivers, candidate non-coding drivers were more enriched in the subclonal category, suggesting that candidate non-coding drivers may contribute more to the subclonal diversification (Extended Data Fig. 7c). In the CLCA, the earliest events were PPP1R12B 3′ UTR mutation and 17p loss, followed by mutations in TP53, ARID2 and the ADH1B 3′ UTR (Extended Data Fig. 7d). By contrast, TP53 mutation was found to be the earliest mutational event in the PCAWG8. Notably, TERT promoter mutations were among the latest events, which was distinct to the observation that TERT promoter was an early event in HCC in European individuals10. These results revealed the distinct evolutionary history of the Chinese CLCA HCC cohort and highlighted the early and pervasive contributions of non-coding mutations during HCC progression. Moreover, the SBS signatures related to tobacco, aflatoxin and AA exposure (SBS_H10, SBS_H3 and SBS_H2), as well as the previously undescribed signature SBS_H8, tended to occur early across all cases (Extended Data Fig. 7e), consistent with that shown in Extended Data Fig. 4h. Furthermore, stratification based on cluster V (SBS_H8), alcohol and smoking revealed distinct evolutionary histories associated with aetiology (Extended Data Fig. 7f and Supplementary Fig. 2). Notably, FGA mutations were among the earliest drivers in patients in cluster V, patients who drink alcohol and patients who smoke.
Metabolic dysregulation
Signalling pathway analysis revealed the higher contributions of non-coding mutations compared with coding mutations in RTK–RAS–MAPK (22.1% versus 6.5%), telomere maintenance (34% versus 1.4%) and liver metabolism (23.1% versus 18.2%), respectively (Extended Data Fig. 8, Supplementary Table 6 and Supplementary Note 7). Particularly, for liver metabolism, a total of 15 potential driver genes was included. These alterations affected various metabolic programs, including hepatic metabolism (APOB, ALB and HNF1A), oxidative stress (KEAP1 and NFE2L2), urea metabolism (CPS1), alcohol metabolism (ADH1B and ADH4), fatty acid metabolism (SERPINA1 and SERBP1) and hypoxia (ARNT). FGA in the JAK–STAT pathway also has a role in hepatic metabolism. Given that the liver is a key metabolic organ and metabolism dysregulation is an important feature of liver cancer20,28, this result underlined the necessity of weighting the contribution of non-coding alterations to investigate the metabolic status of HCC.
KCNJ12 and PPP1R12B
To investigate whether the candidate non-coding drivers have tumorigenic functions, we selected three representative drivers to perform functional assays, including KCNJ12 (potassium inwardly rectifying channel subfamily J member 12), PPP1R12B (protein phosphatase 1 regulatory subunit 12B) (Extended Data Fig. 9, Supplementary Table 7 and Supplementary Note 8) and FGA (Fig. 5, Extended Data Fig. 10 and Supplementary Figs. 3–5). PPP1R12B is one of the earliest driver events, whereas KCNJ12 is one of the latest driver events during the evolutionary history of HCC. Low expression of PPP1R12B significantly enhanced tumour migration, invasion, self-renewal and cell proliferation (Extended Data Fig. 9a). Using the prime editing technology, we showed that point mutations of PPP1R12B identified in the CLCA lead to lower mRNA expression and were enough to cause phenotypic changes (Extended Data Fig. 9b–e). KCNJ12 disruption significantly impaired tumour migration, invasion, self-renewal and cell proliferation (Extended Data Fig. 9f). Point mutations in KCNJ12 lead to a higher level of mRNA expression and subsequent phenotypic changes (Extended Data Fig. 9g–j). These data validated that PPP1R12B and KCNJ12 are non-coding drivers of HCC.
a,b, FGA expression between altered and WT tumours (a) and between paired tumours and normal tissues (b). c,d, FGA protein in paired tumour and normal samples was compared using western blot (WB; c) and immunohistochemistry (IHC; d) analysis. e, Sanger sequencing plots of edited sites in the FGA coding region. f, Quantitative PCR with reverse transcription (RT–qPCR) analysis of FGA mRNA across HepG2 WT and mutated cell lines. n = 3 per group. g, Western blot analysis of FGA. h,i, Comparison of the proliferation (h), and migration, invasion and self-renewal (i) abilities across FGA-edited cell lines. Each assay was repeated three times independently and representative images are shown. For i, scale bars, 100 μm (top and middle) and 3 mm (bottom). j, In vivo cell proliferation assay comparing xenograft tumours of shCtrl (n = 6) and shFGA (n = 7) PLC/PRF/5 cells. Growth curves are shown. k, Representative haematoxylin and eosin (H&E) and immunohistochemistry staining of tumour samples in j. Scale bars, 200 μm (main images) and 25 μm (magnified images). l, The subcellular localization of pTYK2 and pSTAT3. GAPDH (cytoplasmic reference) and lamin A/C (nuclear reference). m, The IL-6 concentration in the supernatant. n = 3 per group. n, Two-tailed Pearson correlation analysis of FGA protein and IL-6 concentration (n = 71). For all panels, n denotes biologically independent samples. For the box plots in a–c, the centre line shows the median, the box limits indicate the upper and lower quartiles, and the whiskers extend to 1.5× the interquartile range; data beyond the whiskers are outlying points. For f, h, j and m, data are mean ± s.e.m. Statistical analysis was performed using two-sided Student’s t-tests (a, f, i and m), two-sided paired t-tests (b–d) and two-way analysis of variance (h and j). Gel source data are provided in Supplementary Figs. 3–5.
FGA dysfunction promotes HCC
Next, we investigated the biological functions of a candidate driver, FGA, which was determined independently as both a candidate coding and non-coding driver (Fig. 5 and Extended Data Fig. 10a). In the CLCA, FGA alterations, including point mutations, loss of heterozygosity and copy-number loss could all result in reduced expression level (Fig. 5a). Meanwhile, the mRNA and protein levels of FGA were lower in tumours compared with the levels in normal tissues (Fig. 5b–d and Extended Data Fig. 10b–d). Furthermore, the rate of biallelic inactivation for FGA was comparable to other recurrently mutated tumour suppressor genes of HCC in the CLCA (Supplementary Table 1). We therefore speculated that FGA is a tumour suppressor gene and explored the potential role of FGA dysfunction in HCC progression.
Induction of FGA point mutations leads to lower mRNA and protein expression and enhanced tumour progression (Fig. 5e–i and Extended Data Fig. 10e–h). Consistent phenotypes were confirmed in FGA-disrupted cell lines (Extended Data Fig. 10i,j). Furthermore, an in vivo assay by subcutaneous injection of short hairpin RNA against FGA (shFGA) cells into BALB/c nude mice resulted in larger and more aggressive tumours in comparison to those of mice injected with shCtrl cells (Fig. 5j,k and Extended Data Fig. 10k). Phosphorylated tyrosine kinase 2 (pTYK2) and its target protein signal transducer and activator of transcription 3 (STAT3, Tyr705) were identified as the top downstream signals of FGA (Extended Data Fig. 10l–n). We also found that pTYK2 accumulated more in the cytoplasm than in the nucleus (Fig. 5l). A specific inhibitor of pTYK2 (BMS-986165), rather than AKT inhibitors, attenuated the migration ability of shFGA cells (Extended Data Fig. 10o). These results suggested that FGA dysfunction might not activate AKT signalling in HCC. We further checked the expression of interleukin-6 (IL-6), a downstream signal of STAT3. The levels of IL6 mRNA and cellular supernatant IL-6 protein were significantly higher in shFGA cells compared with in shCtrl cells (Fig. 5m and Extended Data Fig. 10p,q). Significant negative correlations between FGA and TYK2 phosphorylation, as well as between FGA and IL-6 concentration, were confirmed in an independent HCC cohort (Fig. 5n and Extended Data Fig. 10r). Taken together, our results support that FGA is a tumour suppressor and FGA mutations could promote hepatocarcinogenesis by activating the TYK2–STAT3–IL6 circuit, which could be a potential target for HCC intervention and clinical treatment (Extended Data Fig. 10s).
Discussion
Here we depict a comprehensive whole-genome landscape of HBV-enriched HCC in Chinese individuals. Our high-depth WGS data enabled the identification of previously undescribed candidate non-coding drivers, mutational signatures and subclonal catastrophic events, and the pervasive contribution of non-coding events during HCC evolution. Many of our findings, including the SBS_H8 signature, HBV-ecDNA and distinct aetiology-related evolutionary histories, were highly dependent on the differences between tumours of Chinese and non-Chinese individuals with HCC. These findings shed light on the genomic alterations and processes that are enriched in the tumours of Chinese individuals with HCC. On the other hand, many potential driver events, including candidate driver genes, mutational processes and clustered alterations were shared among our CLCA cohort, the PCAWG-HCC and TCGA-HCC cohort, suggesting universal processes of HCC pathogenesis. In this regard, our findings of previously undescribed non-coding candidates, signatures related to AA and aflatoxin, and subclonal clustered alterations are largely due to the higher depth of the CLCA compared with that of other HCC WGS studies (around 30–40×). These findings should therefore also apply to other HCC cohorts. Notably, 28 non-coding drivers identified in our cohort were previously unreported for HCC, suggesting that our understanding of HCC genome is still very limited.
Although the PCAWG project has characterized 81 mutational signatures across human cancers6, we were able to identify five additional previously undescribed signatures in the CLCA cohort. This result suggested that Chinese patients with HCC have a distinct mutational background in comparison to the members of the cohorts of Japanese and European individuals with HCC. Although SBS_H8 is distinct from AA-related SBS_H2, significant co-occurrence between SBS_H8 and SBS_H2 across the CLCA suggested that the underlying aetiological factors might often co-exist. Future experiments are needed to identify the aetiological factors of SBS_H8.
The high-depth data enabled us to accurately determine the clonal composition of 494 tumours, resulting in the identification of a series of subclonal events. Five out of eight non-coding drivers showed significant enrichments of subclonal mutations. Mutational signatures also exhibited clonality preference, providing important clues for the relative timing of diverse underlying aetiological factors. The identification of subclonal kataegis, chromothripsis and chromoplexy showed that these catastrophic genomic alterations could occur with variable timing during HCC evolution, consistent with the reported combined punctuated and gradual clonal evolution in HCC29. Furthermore, multiple non-coding drivers were mapped to the evolutionary history of CLCA tumours, while the PCAWG reports only one non-coding driver. Our results reconstructed a high-resolution evolutionary history for HCC.
HBV integration has been extensively reported in the HBV-positive tumours of Chinese patients with liver cancer, with hotspots identified in TERT and KMT2B12,30. However, the manner in which these integrations localize in the genome has not been comprehensively assessed. Here we showed that these HBV integrations could be cyclized as ecDNAs. ecDNA amplifications lead to higher levels of oncogene transcription in comparison to copy-number-matched linear DNA21 and they are characterized by enhanced chromatin accessibility31. We identified HBV–oncogene–ecDNA structures, and observed consistent elevated copy numbers and gene expression of HBV together with targeted oncogenes. These results revealed a mechanism of HBV integration in HCC tumorigenesis.
We report a comprehensive genomic landscape of HCC in Chinese individuals covering multiple classes of somatic alterations. How these different genetic alterations cooperate with the diverse immune and stromal cell types in the tumour microenvironment32 is worth in-depth investigation. Collectively, our CLCA study is a valuable resource that provides important biological insights into HCC carcinogenesis and clinical implications to HCC diagnosis and treatment.
Methods
Patient cohort of CLCA
Patients with HCC were enrolled from Eastern Hepatobiliary Surgery Hospital and Shanghai Zhongshan Hospital during 2017–2020. No patients received any preoperative anti-cancer treatment. Each specimen was diagnosed by two senior pathologists. Patients with tissue samples that had sufficient and good-quality DNA were selected. In total, samples from 494 patients with HCC were processed for sequencing analysis, including WGS (n = 494) and RNA-seq (n = 239). Since this is an observational study, no statitistical methods are used to predetermine sample size and no randomization is performed. The study is not an intervention study and therefore blinding is not required. Detailed clinical information is summarized in Supplementary Table 1. DNA from primary tumours and matched peripheral blood lymphocytes was obtained. The study protocol was reviewed and approved by the institutional review board at Eastern Hepatobiliary Surgery Hospital and Shanghai Zhongshan Hospital. This study was performed in accordance with the principles of the Declaration of Helsinki. All of the participants provided written informed consent. All of the samples were anonymously coded in accordance with local ethical guidelines. All research participants consent to the publication of research results.
Cell lines
For the functional validation of three candidate drivers, the human liver cancer cell lines PLC/PRF/5, PVTT, HepG2, Huh7, SNU387 and SNU182, and the normal liver cell line HHL5 were obtained from Shanghai Cell Bank of the Chinese Academy of Sciences. PLC/PRF/5, PVTT, HepG2 and Huh7 cells were cultured in high-d-glucose Dulbecco’s modified Eagle medium (DMEM, Gibco); and SNU387, SNU182 and HHL5 cells in RPMI 1640 medium (basal medium) containing 10% fetal bovine serum (FBS, Gibco), supplemented with 100 U ml−1 penicillin and 100 μg ml−1 streptomycin.
For the validation of AA-related mutational signatures, MCF-10A and HepG2 cells were obtained from the American Type Culture Collection (ATCC). HepG2 cells were cultured as described above. MCF-10A cells were cultured in DMEM/F12 medium supplemented with 10% FBS, 10 ng ml−1 insulin, 20 ng ml−1 EGF, 0.5 µg ml−1 hydrocortisone, 50 ng µl−1 penicillin and 50 U ml−1 streptomycin. All of the cell lines used in this study were authenticated by applying short-tandem-repeat DNA profiling, and were tested to be mycoplasma negative. All of the cell lines were maintained at 37 °C in a humidified incubator with an atmosphere containing 5% CO2.
WGS
Fresh frozen tumour tissues and matched peripheral blood were collected from each patient. DNA was isolated using the DNeasy Blood & Tissue Kit (Qiagen). RNA was extracted using the RNeasy Mini Kit (Qiagen). The DNA concentration was measured using Qubit 3.0 (Invitrogen). The size of the DNA was checked using the Fragment Analyzer (Advanced Analytical Technologies). DNA (200 ng to 1 μg) was sheared into fragments of approximately 300 bp using the Covaris S2 (Covaris) ultrasonicator. The library was constructed using the NEBNext Ultra DNA Library Prep Kit for Illumina (New England Biolabs) according to the manufacturer’s protocol. The library (2 × 150 bp paired-end reads) was quality-checked and sequenced on the Illumina NovaSeq (Illumina) system.
Mutation calling
Raw sequencing reads were processed for quality control by trimming adapter sequences and removing poly(N) and low-quality reads, after which they were preprocessed by FASTP (v.0.13.1) using the following parameters: “--cut_by_quality3 -l 50 --correction -g -x”. The FASTQ files were aligned to the human reference genome (hg19/GRCh37) by Burrows-Wheeler Aligner (BWA, v.0.7.12). Sambamba (v.0.6.8) was used to process PCR duplicates for mapped BAM files. Somatic mutations, including single-nucleotide variants (SNVs) and small insertions and deletions (indels), were called using two methods—Mutect2 (v.4.0.11.0)33 and Strelka2 (v.2.8.4)34.
For Mutect2, a panel of normals (PON) file was first created and somatic mutations were called by comparing each tumour sample with its matched non-tumour counterpart and the PON file. We filtered any mutations with a ‘fragment_length’, ‘mapping_quality’, ‘strand_artifact’, ‘base_quality’ or ‘read_position’. We selected mutations covered by ≥20 reads in the tumour and 10 reads in the normal samples, and excluded mutations belonging to the ENCODE Data Analysis Consortium blacklisted regions. For Strelka2, somatic mutations were called with the flag ‘PASS’. We added an additional quality filter to tighten filtering for low allelic frequency variants: quality score × allele frequency > 1.3. We filtered any variant that was supported by three or more reads in the reference sample in at least three patients. We also filtered indels that were three bases or longer where there was a PON-filtered indel of three bases or longer within ten bases in the same sample. The intersection of the Mutect2 and Strelka results was used as the final set of somatic mutations.
Identification of candidate drivers
We combined P values obtained from independent methods of driver discovery using the empirical Browns method as described in the PCAWG study of non-coding drivers5. Three methods of driver discovery were used for coding regions: MutSigCV35, dndsCV36 and OncodriveFML37. We explored potential non-coding drivers by combining four methods: MutSigCV-NC38, NBR5, ActiveDriverWGS39 and OncodriveFML37. All drivers were manually checked to filter false-positive ‘driver’ loci caused by the sequencing and mapping artefacts, inaccurate background models or local increases in mutations due to mutational processes that were unaccounted for, as previously reported5.
dN/dS analysis
The dN/dS is the ratio between the rates of nonsynonymous and synonymous substitutions, and is used for assessing selection in cancer genomes as described previously17,18. In brief, dN/dS ratios can be calculated for different groups of mutations, such as clonal and subclonal mutations in known cancer genes, yielding insights about the density of driver mutations in each group of mutations. Using the dndscv R package36, dN/dS analysis was run on the clonal and subclonal mutations. A dN/dS ratio of more than 1 indicates positive selection, whereas smaller ratios characterize negative selection, and dN/dS ≈ 1 points toward neutral evolutionary dynamics.
TERT promoter mutation
To double check TERT promoter mutations, we performed targeted sequencing of TERT promoter mutations on tumour samples. The library was constructed by two rounds of PCR amplification. The first round of PCR used a barcoded primer targeting the TERT promoter, which yielded a product of 239 bp. The second round of PCR uses universal indexed primers, yielding a 333 bp product. The sequencing library was then pooled by mixing the PCR products with the same index but with different barcodes. The library was then processed for quality control and sequenced as described for WGS. The average sequencing depth for the region is 378,535×. Data processing was performed the same as for WGS, except that PCR duplicates were not removed.
Mutational signature extraction and assignment
We used mSigHdp (v.1.1.2)40 and SigProfilerExtractor from SigProfiler bioinformatics tool suite (v.1.1.0)6 to extract SBS, DBS and ID signatures. For SigProfiler signature extraction, 1,000 iterations were performed (nmf_replicates = 1000). We report only signatures supported by both mSigHdp and SigProfiler. A signature was considered to be supported by both programs if (1) the mSigHdp-extracted signature has a cosine similarity ≥ 0.90 with a SigProfiler-extracted signature or (2) the mSigHdp-extracted signature can be reconstructed by multiple SigProfiler-extracted signatures (reconstruction cosine similarity ≥ 0.90).
Mutational signature assignment was performed using mSigAct::MAPAssignActivity (v.2.2.3). The prior proportion of each mutational signature was estimated based on the preliminary assignment by mSigHdp. We then performed ‘Ward.D’ hierarchical clustering on the Euclidean distances between signature assignments. For simplification, we combined SBS_H1, SBS_H4, SBS_H14, SBS_H16 and SBS_H17, which were similar to or splits of ageing-related COSMIC signatures SBS1, SBS5 and SBS40, as the ‘Ageing’ SBS signature.
Comparison of extracted signatures to COSMICv3.2 signatures
An extracted signature was confirmed as a known signature if (1) it was similar to a COSMICv3.2 signature (cosine similarity ≥ 0.90); (2) it could be reconstructed by multiple COSMICv3.2 signatures (reconstruction cosine similarity ≥ 0.90); or (3) it could be reconstructed into a COSMICv3.2 signature by combining it with other extracted signatures (reconstruction cosine similarity ≥ 0.90). Steps (2) and step (3) were evaluated using mSigAct::OptimizeExposureQP. Pentanucleotide context analysis includes 2 bp before and after the mutation. As for SBS_H8, the low overall cosine similarity of 0.71 between SBS_H8 and SBS22 lead us to further perform a comparison of pentanucleotide context. The low pentanucleotide cosine similarity of 0.61 further revealed substantial differences between SBS_H8 and COSMIC SBS22. Specifically, T>A mutations of SBS_H8 were enriched in the NCxGG context, while that of SBS22 had a rather dispersed NCxGN context.
Cell culture to validate the new mutational signatures
Exposure of HepG2 and MCF-10A cells was performed as previously described41. In brief, HepG2 cells were exposed to 20 µM of aristolochic acid 1 (AA1, A5512, Sigma-Aldrich) for 2 months, whereas MCF-10A cells were exposed to 20 µM or 40 µM AA for the same length of time. After 2 weeks of recovery and expansion, single-cell cloning was performed using flow cytometry. Random clones were selected and expanded for DNA isolation and WGS. For MCF-10A cells, we sequenced two clones from the cells exposed to 20 µM AA (clones 1 and 2), as well as one clone from the cells exposed to 40 µM AA (clone 3). For HepG2 cells, we sequenced all three clones from the cells exposed to 20 µM AA.
CNA
Sequenza (v.2.1.1) was used to call CNAs, taking both ploidy and cellularity into account42. In brief, we used BAM files of tumour and paired normal samples as an input to calculate the depth ratio, which was normalized using the GC content bias and data ratio. To acquire segmented copy numbers and estimate cellularity and ploidy, the following parameters were used: breaks.method = ‘full’, gamma = 40, kmin = 5, gamma.pcf = 200, kmin.pcf = 200. For each tumour sample, the copy numbers of segments were divided by ploidy after log2 transformation. After filtering out segments smaller than 500 kb, copy-number states were determined for each segment. Copy-number gains and losses were defined as at least one copy more and one copy less than the estimated ploidy, respectively.
PURPLE (PURity & PLoidy Estimator; v.2.34) was also performed on paired tumour-normal WGS data as described previously43. There are five key steps in the PURPLE pipeline, as follows: (1) calculate the tumour B-allele frequency at high-confidence heterozygous germline loci; (2) determine read-depth ratios for tumour and reference genomes; (3) segmentation; (4) purity fitting; (5) smoothing. A number of rules were further applied to merge adjacent regions to create a smooth copy-number profile. GISTIC2.0 (Genomic Identification of Significant Targets in Cancer v.2.0.23) was used to identify focal gain and loss regions44.
SVs
SVs were called using LUMPY (v.0.2.13) using the default parameters45. LUMPY simultaneously integrates multiple SV detection signals during SV discovery. Both read-pair and split-read signals were considered within the LUMPY framework, achieving a relatively high detection sensitivity for SVs. The detected SVs were further used for analysis of clustered mutational processes, including kataegis, chromothripsis and chromoplexy.
Kataegis
Kataegis is a focal hypermutation process that leads to locally clustered point mutations25. Kataegis events are defined as genomic segments containing six or more consecutive mutations with an average intermutation distance of less than or equal to 100 bp. Rainfall plots containing kataegis were plotted by the rainfallPlot function with detectChangePoints set to TRUE from the R package Maftools (v.2.6.05)46.
Chromothripsis
Chromothripsis is characterized by massive genomic rearrangements exhibiting oscillations between two copy-number states23. Chromothripsis was inferred using the R package ShatterSeek (v.0.4)26. In brief, it first uses intrachromosomal SVs to detect clusters of interleaved rearrangements. Next, it evaluates a set of statistical criteria in each of these regions. The output consists of a data frame reporting the value for the statistical criteria used and additional information for each chromosome. Candidate chromothripsis regions were visually inspected with the local SVs and CN profiles. For the minimum number of oscillating CN segments, we used two thresholds: high-confidence calls display oscillations between two states in at least seven adjacent segments, whereas low-confidence calls involve between four and six segments.
Chromoplexy
Chromoplexy results from several simultaneous double-stranded DNA breaks in several chromosomes that are rejoined incorrectly, leading to balanced chains of rearrangements7,24. Chromoplexy was inferred by ChainFinder (v.1.0.1), an algorithm for identifying complex sets of DNA rearrangements and deletions in cancer genomes that may reflect coordinate chromosomal alterations24. In brief, ChainFinder first models the expected chromosomal distribution of breakpoints from independently arising rearrangements. The algorithm then profiles user-provided copy-number and SV data for sets of rearrangements and associated gene deletions that are unlikely to have arisen independently based on their deviation from the predicted distribution.
HBV integration
We first aligned all reads against a comprehensive list of HBV virus reference sequences as described previously (n = 73)14. We next searched for human–virus chimeric reads, where one end or one part of the read was mapped to the human genome, while the other end or the left part of the read was mapped to the viral reference genome, because these reads indicate HBV integration into the human genome. Adjacent or overlapping chimeric reads (within 500 bp) aligning to the human and viral genomes in the same orientation were merged to make clusters. Clusters with at least two chimeric reads were retained. The integration sites were then compared to RefSeq gene boundaries to find genes that were directly disrupted by HBV integration (overlapping) or potentially affected by integration (within 15 kb of integration sites). HBV fusion was also detected using the RNA-seq data with STAR-Fusion47.
Detection of ecDNA
ecDNA-based amplification has been recognised as a way for tumour cells to increase the copy number of oncogenes22,48,49. AmpliconArchitect (v.1.3.r2) was used to detect ecDNA22. In brief, aligned reads of regions with CN greater than five were used as seeds. The default parameters were used. Given mapped reads, AmpliconArchitect automatically searches for other intervals participating in the amplicon, and then uses a combination of CNV and SV analysis. AmpliconArchitect uses structural variant signatures (for example, discordant paired-end reads and CNV boundaries) to partition all intervals into segments and build an amplicon graph. It assigns CNs to the segments by optimizing a balanced flow on the graph. We then used the AmpliconArchitect-derived breakpoint graph to classify amplicons into four categories using AmpliconClassifier (v.0.2.5): (1) circular amplification; (2) breakage–fusion–bridge amplification; (3) heavily rearranged amplification; and (4) linear amplification as described21. Circular amplicons were considered to be ecDNA.
Validating ecDNA with CIRCLE-seq
CIRCLE-seq is a sequencing library enrichment approach optimized for circular DNA detection50,51 and was performed on selected cases. A detailed protocol for circular DNA isolation is available on the Nature Protocol Exchange server (https://doi.org/10.1038/protex.2019.006). Amplified circular DNA was sheared to an average fragment size of 150–200 bp using the S220 focused ultrasonicator (Covaris). Libraries for next-generation sequencing were prepared using the NEBNext Ultra DNA Library Kit for Illumina according to the manufacturer’s protocol (New England Biolabs). Sequencing data generated by CIRCLE-seq were aligned and processed. The aligned BAM files were then analysed in two ways. First, all read pairs and split reads containing any outward-facing read orientation, indicating potential circles, were placed into a new BAM file. Second, genomic segments enriched for signal over background were detected in the ‘all reads’ BAM file using variable-width windows from Homer v.4.11 findPeaks, and the edges of these enriched regions were intersected with the ‘circle only’ BAM file to quantify the number of circle-supporting reads. To determine the thresholds for significance of real circles versus background noise, matched WGS data were used to determine the background distribution of circle-oriented reads in non-circle-enriched regions that were matched for length and nucleotide composition. An empirical P value of 0.01 was used to filter putative circles, and regions passing this filter were then used for downstream analysis.
Inferring clonality and evolutionary history
The evolutionary history of our CLCA cohort was determined as previously described8. In basic terms, clonal mutations occurred before the emergence of the most-recent common ancestor, whereas subclonal mutations occurred after this event. In regions with copy-number gains, molecular time can be further divided according to whether mutations preceded the copy-number gain (and were themselves duplicated) or occurred after the gain and were therefore present on only one chromosomal copy. In brief, the variant allele frequencies (VAFs) of somatic point mutations cluster around the values imposed by the purity of the sample and local copy-number states. On the basis of this information, subclonal populations were identified, the timing of copy number gains and point mutations was inferred, and the relative timing of somatic driver events was deduced. We next inferred the timing of mutational signatures. We inferred the mutational history of our CLCA cohort by integrating these timing data across 494 patients. We also divided these patients according to aetiological factors, such as smoking and drinking, and compared the evolutionary history of the different groups. Key packages used for timing analysis, including PyClone (v.0.13.1), MutationTimeR (v.0.99.3), and PhylogicNDT (v.1.0), are available at the PCAWG GitHub repository (https://github.com/PCAWG-11/Evolution).
RNA-seq
RNA in the tumour samples was extracted using the RNeasy Mini Kit (Qiagen). The DNA and RNA concentration was measured using Qubit 3.0 (Invitrogen). The size of RNA was checked using Fragment Analyzer (Advanced Analytical Technologies). RNA-seq libraries were constructed using the TruSeq mRNA Library Prep Kit (Illumina) according to the manufacturer’s protocol. The library (2 × 150 bp paired-end reads) was then quality checked and sequenced using the Illumina NovaSeq (Illumina) system. Qualified reads were obtained after removing raw reads with adapters or of low quality and then aligned to the human genome (hg19) using STAR (v.2.7.3c)52. The transcripts per million (TPM) values and gene count values were computed using RSEM (v.1.3.3). Fusion genes were detected using STAR-Fusion47.
Stable cell line construction
Three representative candidate drivers were selected for functional validation, including 3′ UTR PPP1R12B, KCNJ12 promoter and FGA. 3′ UTR PPP1R12B was among the earliest candidate driver events during HCC evolution while the KCNJ12 promoter was among the latest driver events. Particularly, FGA, was determined independently as both candidate coding and non-coding drivers (3′ UTR). Knockdown by short hairpin RNA (shRNA) or knockout by short guide RNA (sgRNA) for the three drivers were constructed on a total of seven human cell lines, including six liver cancer cell lines (PLC/PRF/5, PVTT, SNU387, SNU182, Huh7 and HepG2) and one normal liver cell line (HHL5). The lentiviruses of shRNA or sgRNA targeting the above three genes were obtained (Supplementary Table 7) and transfected into cell lines as indicated.
For PPP1R12B, disrupted cell lines were constructed using two independent shRNAs (1 and 2) in PLC/PRF/5, PVTT, SNU387 and HHL5 cells, and by two independent sgRNAs (1 and 2) in HepG2 and Huh7 cells. For KCNJ12, disrupted cell lines were constructed by two independent shRNAs (1 and 2) in SNU182 and HepG2 cells, and by two independent sgRNAs (1 and 2) in PLC/PRF/5, PVTT, SNU387 and HHL5 cells. For FGA, disrupted cell lines were constructed by two independent shRNAs (1 and 2) in PLC/PRF/5, PVTT, SNU387 and SNU182 cells, and by two independent sgRNAs (1 and 2) in Huh7 cells. Scramble shRNA was used as a control (shCtrl).
We failed to knock in the detected non-coding 3′ UTR mutations of FGA using either Prime Editing technology or other base editors, such as cytosine base editor (CBE) or the adenine base editor (ABE). To confirm the functional role of FGA 3′ UTR mutations, we first knocked out endogenous FGA by sgRNA in the HepG2 cell line and then induced ectopic stable expression of mutant 3′ UTR with the wild type as a control (Extended Data Fig. 10e). Overexpression lentiviruses containing FGA 3′ UTR mutation with the wild type as a control were constructed by Ubigene Biosciences and were ectopically stably expressed in HepG2 single-cell clones without endogenous FGA. In total, 2.5 × 105 cells were plated into six-well plates, incubated overnight and transfected with lentiviral particles (multiplicity of infection of 10) the next day. At 12–24 h after transduction, the medium was replaced with complete culture medium for 72 h, and the stable knockdown or knockout cell lines were sorted by flow cytometry or puromycin.
Prime editing
Endogenous point mutants (FGA, PPP1R12B and KCNJ12) were introduced using the Prime Editing (PE) technology in the HepG2 cell line. In brief, plasmid expression of prime editing guide RNAs (pegRNAs) or nicking sgRNAs were cloned using Golden Gate assembly as previously described53. pegRNA was cloned into pU6-tevopreq1-GG-acceptor plasmid (Addgene) with an inserted EF1α promoter and puromycin-resistance cassette. Nicking sgRNA used for PE3 or PE3b was cloned into BPK1520 (Addgene). In total, 3 × 105 cells were plated into a 24-well plate overnight, and transfected at approximately 80–90% confluency with 1,000 ng pCMV-PEmax-P2A-BSD plasmid (Addgene), 500 ng pCMV-hMLH1dn plasmid (Addgene), 333 ng pegRNA plasmid and 111 ng nicking sgRNA plasmid by Lipofectamine 3000 (Invitrogen) according to the manufacturer’s instructions. Cells were cultured and sorted by puromycin and blasticidin. Genomic DNA from edited clones was extracted, and the targeting region was amplified by PCR and sequenced on ABI 3730XL (Thermo Fisher Scientific). A list of all of the pegRNAs, nicking sgRNAs and primer sequences is provided in Supplementary Table 7b,c.
RT–qPCR analysis
Total RNA from HCC cell lines was isolated using TRIzol Reagent (Invitrogen) according to the standard instructions. RNA was reverse transcribed into first-strand cDNA using 1 μl of random hexamers (Bio-Light Biotech), 1.25 μl Recombinant RNasin Ribonuclease Inhibitor (Promega), 1 μl 4 × dNTP Mixture (Bio-Light Biotech), 1 μl M-MLV reverse transcriptase (Promega) and 5 μl M-MLV RT 5× buffer (Promega). qPCR was performed using the ChamQ SYBR Colour qPCR Master Mix (Vazyme) on the LightCycler 96 PCR platform (Roche). A list of the sequences of the specific RT–qPCR primers is provided in Supplementary Table 7c. The cycling conditions were as follows: 95 °C for 10 min, 45 cycles of 95 °C for 10 s, and 60 °C for 30 s. The results were normalized to ACTB (encoding β-actin) mRNA expression and analysed using the \({2}^{-\Delta \Delta {C}_{{\rm{t}}}}\) method.
Western blotting
Total protein from frozen tissue samples and cell lines was lysed in RIPA lysis buffer (Strong) (Yesen) in the presence of 1% protease inhibitors and phosphatase inhibitors (Yesen). The concentration of protein was assessed using the Pierce BCA Protein Assay Kit (Thermo Fisher Scientific) according to the manufacturer’s protocol. Equal amounts of total protein were separated by 8% SDS–PAGE and transferred onto preactivated poly vinylidene fluoride membranes (Millipore). The blots were incubated with the appropriate primary antibodies (4 °C, overnight) against β-actin (AC004, AMC0001, 1:5,000; ABclonal), GAPDH (AC033, AMC0062, 1:5,000; ABclonal), TYK2 (9312, 1:1,000; Cell Signaling Technology), phosphorylated-TYK2 (Tyr1054/1055) (D7T8A) (68790, D7T8A, 1:1,000; Cell Signaling Technology), phosphorylated-STAT3 (Tyr705) (D3A7) XP (9145, D3A7, 1:2,000; Cell Signaling Technology), lamin A/C (4C11) (4777, 4C11, 1:2,000; Cell Signaling Technology), fibrinogen-α (C-7) (sc-398806, C-7, 1:500; Santa Cruz Biotechnology), STAT3 (60199-1-Ig, 3G2D12, 1:2,000; Proteintech). Next, the bands were incubated with HRP-conjugated goat anti-rabbit IgG (H+L) (SA00001-2, 1:5,000; Proteintech) and goat anti-mouse IgG (H+L) (SA00001-1, 1:5,000; Proteintech) or fluorescently labelled IRDye 800CW goat anti-rabbit IgG (H+L) (926-32211, 1:20,000; LI-COR) and IRDye 800CW goat anti-mouse IgG (H+L) (926-32210, 1:20,000; LI-COR) secondary antibodies (room temperature, 2 h) listed in Supplementary Table 7d. Immunoreactive bands were detected using the Touch Imager XLi system (e-BLOT Life Science) or Odyssey Sa Infrared Imaging System (LI-COR Biosciences). For the expression of different proteins in the same blots, partly blotted membranes were incubated with western blot fast stripping buffer (EpiZyme) followed by several washes and treated as mentioned above. The band intensity of western blots was assessed using ImageJ (v1.53a). β-Actin, GAPDH and lamin A/C were used as references.
Subcellular fractionation
Nuclear and cytoplasmic fractions of cells were prepared using the Nuclear and Cytoplasmic Extraction Reagents Kit (Beyotime), as well as protease inhibitors, phosphatase inhibitors and phenylmethanesulfonyl fluoride. In brief, cells were washed with precooled 1× phosphate-buffered saline (PBS), after which the cytoplasmic protein samples were collected using Cytoplasmic Protein Extraction Reagent, which disrupted the plasma membranes, leaving the nuclear membranes intact. Nuclear proteins were isolated from the remaining pellet using nuclear protein extraction reagent, followed by western blotting.
Cell proliferation assay
In vitro cell proliferation was assessed using the Cell Counting Kit-8 (CCK8, DOJinDO) according to the manufacturer’s protocol. In brief, the cells were seeded in 96-well plates (0.75–2 × 103 cells per well). At the indicated time, 10 μl of CCK-8 solution was added to each well, and the plates were incubated in the dark at 37 °C for 1–2 h. The spectrometric absorbance of each well at 450 nm was measured using the Synergy Neo microplate reader (BioTek). Data were normalized to day 0, and the results are presented as the fold change over the control samples.
Colony-formation assay
The self-renewal ability of cells was determined using a colony-formation assay. Cells were plated in six-well plates at a density of 1.5–4 × 103 cells per well and cultured in complete medium at 37 °C for 9–21 days. The medium was replaced every 3 days. After the culture period, the cells were fixed with 4% paraformaldehyde for 15 min and stained with 0.1% (w/v) crystal violet for 15 min. Cell confluence in each well was quantified, and the results are presented as the fold change over the control samples.
Cell migration and invasion assay
Migration and invasion assays were performed as previously described using 8-μm-pore-size Transwell chambers (Greiner Bio-one; Falcon; Costar). For cell migration assays, 2–10 × 104 cells prepared in FBS-free medium were seeded onto the upper chambers, while, the lower chamber was filled with 750 μl conditioned medium containing 10–30% FBS. For cell invasion assays, Matrigel-coated Transwell chambers were purchased from Corning and homemade (the chamber inserts were precoated with appropriate proportion Matrigel (Corning) for approximately 2 h in a 37 °C incubator). Next, cell suspension (2.5–10 × 104 cells) diluted in FBS-free medium was seeded to the upper chamber and conditioned media with 10–30% FBS was added to the bottom chamber of the Transwell. After incubation 24–96 h, cells on the upper surface of the membrane were removed with cotton tips. Cells that attached to the lower surface were fixed in 4% paraformaldehyde and stained with 0.1% crystal violet for 15 min. Excess dye was removed by washing the stained cells with water, after which they were examined using the Olympus IX73 microscope equipped with an DP80 camera. For inhibitor treatment experiment, PLC/PRF/5-shCtrl and PLC/PRF/5-shFGA cells were pretreated with pTYK2 inhibitor (BMS-986165, 10 μM) or two AKT inhibitors (MK-2206, 2 μM; AZD5363, 10 μM), and those inhibitors at the same concentration were added to the top and bottom chambers simultaneously. The results are presented as the fold change over the control samples.
Subcutaneous xenograft
BALB/c nude mice (aged 5–7 weeks) were obtained from GemPharmatech. All of the mice were housed in specific-pathogen-free conditions at an ambient temperature of 20–26 °C and a humidity of 30–70% under a 12 h–12 h light–dark cycle before use. Mice had unrestricted access to regular mouse chow and water. Body-weight-matched mice were randomized for subcutaneous injection into treatment groups. Blinding was not required. We subcutaneously injected 2 × 106 PLC/PRF/5-shCtrl or PLC/PRF/5-shFGA cells within 100 μl of PBS/Matrigel (3:2) into the flanks of nude mice (shCtrl, n = 6; shFGA, n = 7). For the sample size, a minimum of three mice for each group of the PLC/PRF/5-shCtrl and PLC/PRF/5-shFGA cells was required to reach statistical significance. Preliminary subcutaneous xenograft experiments were performed on male and female mice, respectively. Similar trends of shFGA cells resulted in larger and more aggressive tumours in comparison to those of mice injected with shCtrl cells were observed. To exclude the potential confounding factors of aggression and biting in the male groups, only the female groups were retained and recorded. The tumour width (w) and length (l) were measured every 3 days using callipers and the diameter of a single tumour was <2 cm at the time of euthanasia. Tumour volume (V) was calculated individually using the following formula: V = (w2 × l) × 0.52. Tumour tissues were embedded in paraffin wax and cut in slices, followed by immunohistochemistry (IHC). All of the mouse experiments were approved by the Animal Care and Use Committee at Eastern Hepatobiliary Surgery Hospital.
Immunohistochemistry analysis
We collected HCC tissue microarrays (n = 39) and tumour samples from xenograft mouse models, which were fixed in 10% neutral formalin, embedded in paraffin and cut into 3–5 μm sections on charged glass slides. After deparaffinization, rehydration, blocking endogenous peroxidase and heat-induced antigen retrieval, the sections were incubated overnight with primary antibodies at 4 °C including, anti-fibrinogen alpha chain (20645-1-AP, 1:100; Proteintech), anti-phosphorylated-TYK2 (Tyr1054/1055) (D7T8A) (68790, D7T8A, 1:100; Cell Signaling Technology) and anti-Ki-67 (ab15580, 1:500; Abcam). HRP-conjugated anti-rabbit (D-3002; Supervision) secondary antibodies were added for 30 min at 37 °C. The slides were visualized using the Liquid DAB+ Substrate Chromogen System (DAKO) and counterstained with haematoxylin. IHC slides were scanned using the Leica Aperio AT2 system, and images were analysed using Aperio ImageScope (v.12.4.6).
IL-6 concentration
The IL-6 concentration in the cellular supernatant in the PLC/PRF/5 and PVTT cell lines was quantified using the human IL-6 ELISA kit (RayBiotech) according to the manufacturer’s instructions using a Synergy Neo microplate reader (BioTek). The IL-6 concentration in patient tissue was measured using the S-PLEX Human IL-6 Kit (Meso Scale Discovery) according to the manufacturer’s protocol. In brief, all protein samples were prediluted 2–5×. The plates were then assembled, enhanced and read using the 1300 MESO QuickPlex SQ 120MM instrument, which recorded electrochemiluminescence and analysed with the DISCOVERY WORKBENCH Desktop Analysis Software (v.4.0). The concentrations of IL-6 in the samples were interpolated against a standard curve.
FGA expression
To compare the FGA protein level between tumours and matched normal tissues, western blotting and immunohistochemistry were performed on 47 and 39 tumour–normal tissue pairs, respectively. To examine the association among FGA, pTYK2 (Tyr1054) and IL-6 at the protein level, western blots of FGA and pTYK2 (Tyr1054) and electrochemiluminescence signals of IL-6 concentration were assessed in 75 and 71 patients with HCC, respectively. The IL-6 concentration was not available in 4 out of 75 patients. The relative intensity of FGA and pTYK2 were normalized to β-actin.
Phospho-specific protein microarray
The Phospho Explorer Antibody Array (PEX100) was obtained from Full Moon Biosystems and used according to the manufacturer’s protocol. Each of the antibodies printed on the coated glass microscopy slide has two replicates along with multiple positive and negative controls. The phospho-antibody array contained 1,318 site-specific antibody profiles, of which 584 were pairs of phosphoproteins and their unphosphorylated counterparts. Lysates of PLC/PRF/5-shCtrl and PLC/PRF/5-shFGA cells were applied to Phospho Explorer Antibody Arrays, which were applied and analysed by OE Biotechnology. Next, the protein samples were biotinylated and hybridized according to the manufacturer’s protocol. The fluorescence intensity of each antibody spot was obtained using the GenePix 4000B Microarray Scanner (Molecular Devices) and analysed using GenePix Pro v.6.0.
For data analysis, background signals were first removed from all measurements. Second, for each antibody, the respective negative control value was removed from each measurement. Third, if a phosphorylation site did not satisfy the requirements ((1) for each phosphorylated site, two replicates showed the same pattern between shCtrl and shFGA cells; (2) CV < 0.3 for the indicated group with a higher phosphorylation level), that site was considered to be a discrete point set, and was discarded. Next, the phosphorylation ratio between groups was measured using the following formula: phosphorylation ratio = [phosphorylation ratio of the shFGA group (phosphorylation value/unphosphorylation value)]/[phosphorylation ratio of the shCtrl group]. Finally, candidate dysregulated phosphorylated proteins were selected by identifying proteins with a phosphorylation ratio of >1.5 or <0.667 in shFGA cells versus shCtrl cells.
Statistical analysis
Statistical analyses were performed using R (v.3.6.0) and GraphPad Prism (v.9.0). n denotes biologically independent samples unless otherwise specified. The data are presented as the mean ± s.e.m. unless otherwise specified. For box plots in all panels, the centre line shows median, the box limits indicate the upper and lower quartiles, and the whiskers extend to 1.5× the interquartile range, and data beyond the end of the whiskers are outlying points that are plotted individually. All P values were calculated using two-sided analysis. Unpaired Student’s t-tests were used unless otherwise specified. P < 0.05 or Q < 0.1 was considered to be significant.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The raw sequencing data reported in this paper have been deposited at the Genome Sequence Archive in BIG Data Center, Beijing Institute of Genomics (BIG), Chinese Academy of Sciences, under the study accession number PRJCA002666. We also built an interactive website (http://lifeome.net/database/liver) for visualizing and analysing our CLCA data. The data deposited and made public are compliant with the regulations of the Ministry of Science and Technology of China. Other public data used in this study include the human reference genome hg19/GRCh37 (https://ftp.ensembl.org/pub/grch37/), PCAWG data (https://dcc.icgc.org/pcawg/#!), TCGA-HCC data (https://portal.gdc.cancer.gov/projects/TCGA-LIHC) and COSMIC signatures (https://cancer.sanger.ac.uk/signatures/). Source data are provided with this paper.
Code availability
The Linux working environment that we used is packed into a Singularity container file and is available at Zenodo (https://doi.org/10.5281/zenodo.7260221). The detailed codes and instructions for all software have been deposited at GitHub (https://github.com/ChongJenniferZhang/CLCA_WGS).
References
Sung, H. et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 71, 209–249 (2021).
Llovet, J. M. et al. Hepatocellular carcinoma. Nat. Rev. Dis. Primers 7, 6 (2021).
Villanueva, A. Hepatocellular Carcinoma. N. Engl. J. Med. 380, 1450–1462 (2019).
The ICGC/TCGA Pan-Cancer Analysis of Whole Genomes Consortium. Pan-cancer analysis of whole genomes. Nature 578, 82–93 (2020).
Rheinbay, E. et al. Analyses of non-coding somatic drivers in 2,658 cancer whole genomes. Nature 578, 102–111 (2020).
Alexandrov, L. B. et al. The repertoire of mutational signatures in human cancer. Nature 578, 94–101 (2020).
Li, Y. et al. Patterns of somatic structural variation in human cancer genomes. Nature 578, 112–121 (2020).
Gerstung, M. et al. The evolutionary history of 2,658 cancers. Nature 578, 122–128 (2020).
Fujimoto, A. et al. Whole-genome mutational landscape and characterization of noncoding and structural mutations in liver cancer. Nat. Genet. 48, 500–509 (2016).
Letouze, E. et al. Mutational signatures reveal the dynamic interplay of risk factors and cellular processes during liver tumorigenesis. Nat. Commun. 8, 1315 (2017).
Gao, Q. et al. Integrated proteogenomic characterization of HBV-related hepatocellular carcinoma. Cell 179, 561–577 (2019).
Sung, W. K. et al. Genome-wide survey of recurrent HBV integration in hepatocellular carcinoma. Nat. Genet. 44, 765–769 (2012).
Kan, Z. et al. Whole-genome sequencing identifies recurrent mutations in hepatocellular carcinoma. Genome Res. 23, 1422–1433 (2013).
Xue, R. et al. Variable intra-tumor genomic heterogeneity of multiple lesions in patients with hepatocellular carcinoma. Gastroenterology 150, 998–1008 (2016).
Schulze, K. et al. Exome sequencing of hepatocellular carcinomas identifies new mutational signatures and potential therapeutic targets. Nat. Genet. 47, 505–511 (2015).
Imielinski, M., Guo, G. & Meyerson, M. Insertions and deletions target lineage-defining genes in human cancers. Cell 168, 460–472 (2017).
Dentro, S. C. et al. Characterizing genetic intra-tumor heterogeneity across 2,658 human cancer genomes. Cell 184, 2239–2254 (2021).
Martincorena, I. et al. Tumor evolution. High burden and pervasive positive selection of somatic mutations in normal human skin. Science 348, 880–886 (2015).
Tarabichi, M. et al. Neutral tumor evolution? Nat. Genet. 50, 1630–1633 (2018).
Ng, S. W. K. et al. Convergent somatic mutations in metabolism genes in chronic liver disease. Nature 598, 473–478 (2021).
Kim, H. et al. Extrachromosomal DNA is associated with oncogene amplification and poor outcome across multiple cancers. Nat. Genet. 52, 891–897 (2020).
Deshpande, V. et al. Exploring the landscape of focal amplifications in cancer using AmpliconArchitect. Nat. Commun. 10, 392 (2019).
Stephens, P. J. et al. Massive genomic rearrangement acquired in a single catastrophic event during cancer development. Cell 144, 27–40 (2011).
Baca, S. C. et al. Punctuated evolution of prostate cancer genomes. Cell 153, 666–677 (2013).
Nik-Zainal, S. et al. The life history of 21 breast cancers. Cell 149, 994–1007 (2012).
Cortes-Ciriano, I. et al. Comprehensive analysis of chromothripsis in 2,658 human cancers using whole-genome sequencing. Nat. Genet. 52, 331–341 (2020).
Alexandrov, L. B. et al. Signatures of mutational processes in human cancer. Nature 500, 415–421 (2013).
Satriano, L., Lewinska, M., Rodrigues, P. M., Banales, J. M. & Andersen, J. B. Metabolic rearrangements in primary liver cancers: cause and consequences. Nat. Rev. Gastroenterol. Hepatol. 16, 748–766 (2019).
Guo, L. et al. Single-cell DNA sequencing reveals punctuated and gradual clonal evolution in hepatocellular carcinoma. Gastroenterology 162, 238–252 (2022).
Xue, R. et al. Genomic and transcriptomic profiling of combined hepatocellular and intrahepatic cholangiocarcinoma reveals distinct molecular subtypes. Cancer Cell 35, 932–947 (2019).
Wu, S. et al. Circular ecDNA promotes accessible chromatin and high oncogene expression. Nature 575, 699–703 (2019).
Xue, R. et al. Liver tumour immune microenvironment subtypes and neutrophil heterogeneity. Nature 612, 141–147 (2022).
Cibulskis, K. et al. Sensitive detection of somatic point mutations in impure and heterogeneous cancer samples. Nat. Biotechnol. 31, 213–219 (2013).
Kim, S. et al. Strelka2: fast and accurate calling of germline and somatic variants. Nat. Methods 15, 591–594 (2018).
Lawrence, M. S. et al. Mutational heterogeneity in cancer and the search for new cancer-associated genes. Nature 499, 214–218 (2013).
Martincorena, I. et al. Universal patterns of selection in cancer and somatic tissues. Cell 171, 1029–1041 (2017).
Mularoni, L., Sabarinathan, R., Deu-Pons, J., Gonzalez-Perez, A. & López-Bigas, N. OncodriveFML: a general framework to identify coding and non-coding regions with cancer driver mutations. Genome Biol. 17, 128 (2016).
Lawrence, M. S. et al. Discovery and saturation analysis of cancer genes across 21 tumour types. Nature 505, 495–501 (2014).
Zhu, H. et al. Candidate cancer driver mutations in distal regulatory elements and long-range chromatin interaction networks. Mol. Cell 77, 1307–1321 (2020).
Liu, M., Wu, Y., Jiang, N., Boot, A. & Rozen, S. G. mSigHdp: hierarchical Dirichlet process mixture modeling for mutational signature discovery. NAR Genom. Bioinform. 5, lqad005 (2023).
Boot, A. et al. In-depth characterization of the cisplatin mutational signature in human cell lines and in esophageal and liver tumors. Genome Res. 28, 654–665 (2018).
Favero, F. et al. Sequenza: allele-specific copy number and mutation profiles from tumor sequencing data. Ann. Oncol. 26, 64–70 (2015).
Priestley, P. et al. Pan-cancer whole-genome analyses of metastatic solid tumours. Nature 575, 210–216 (2019).
Mermel, C. H. et al. GISTIC2.0 facilitates sensitive and confident localization of the targets of focal somatic copy-number alteration in human cancers. Genome Biol. 12, R41 (2011).
Layer, R. M., Chiang, C., Quinlan, A. R. & Hall, I. M. LUMPY: a probabilistic framework for structural variant discovery. Genome Biol. 15, R84 (2014).
Mayakonda, A., Lin, D.-C., Assenov, Y., Plass, C. & Koeffler, H. P. Maftools: efficient and comprehensive analysis of somatic variants in cancer. Genome Res. 28, 1747–1756 (2018).
Haas, B. J. et al. Accuracy assessment of fusion transcript detection via read-mapping and de novo fusion transcript assembly-based methods. Genome Biol. 20, 213 (2019).
Turner, K. M. et al. Extrachromosomal oncogene amplification drives tumour evolution and genetic heterogeneity. Nature 543, 122–125 (2017).
deCarvalho, A. C. et al. Discordant inheritance of chromosomal and extrachromosomal DNA elements contributes to dynamic disease evolution in glioblastoma. Nat. Genet. 50, 708–717 (2018).
Tsai, S. Q. et al. CIRCLE-seq: a highly sensitive in vitro screen for genome-wide CRISPR–Cas9 nuclease off-targets. Nat. Methods 14, 607–614 (2017).
Koche, R. P. et al. Extrachromosomal circular DNA drives oncogenic genome remodeling in neuroblastoma. Nat. Genet. 52, 29–34 (2020).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Anzalone, A. V. et al. Search-and-replace genome editing without double-strand breaks or donor DNA. Nature 576, 149–157 (2019).
Acknowledgements
We thank D. Li, S. Yin and C. Zhang for their support in gene editing and the members of the Shanghai Key Laboratory of Hepato-biliary Tumour Biology and the Key Laboratory of Signaling Regulation and Targeting Therapy of Liver Cancer (SMMU) for their technical support. This work was supported by the National Natural Science Foundation of China (81988101, T2125002, 82322047, 82241230, U21A20376, 81830054, 82173035, 82141103 and 82341007), the Innovation Program of Shanghai Municipal Education Commission (21JC1406600 and 22140901000), Beijing Natural Science Foundation (Z220014), Beijing Nova Program (20230434854), Program of Shanghai Academic/Technology Research Leader (21XD1404600), the National Key Research and Development Program of China (2022YFC3400902 and 2022YFC2504602), and the New Cornerstone Science Foundation through the XPLORER PRIZE. Figure 1a and Extended Data Fig. 10s were created using BioRender with an academic license.
Author information
Authors and Affiliations
Contributions
L.C., C.Z., R.X., L.W., F.B., S.G.R. and H.W. conceived and designed the project. L.C., Z.L., B.Z., K.L., Y. Zhu, S.Y. and Q.G. collected the clinical samples. C.Z., R.X., M.L., J. Bai, Yin Wang, R.W., A.Y. and Yan Wang analysed the WGS and RNA-seq data. S.G.R., M.L., N.J., C.Z. and R.X. performed mutational signature analysis. L.C., J. Bao, W.W., J.H., S.S., Y. Zhang and M.B. performed functional validation of candidate drivers. R.X., C.Z., J. Bai, L.C. and J.G. designed and built the CLCA website. C.Z., R.W. and N.J. built the Zenodo and GitHub pages. R.X., L.C., C.Z. and J. Bao integrated the sequencing and experimental data, drew the display items and wrote the manuscript. F.B., L.W., D.G., X.W.W., N.Z., H.N., S.G.R. and H.W. provided edits to the manuscript. L.C., L.W., F.B. and H.W. oversaw the ethical guidelines and data regulation. L.C., L.W., F.B., S.G.R. and H.W. supervised the project. All of the authors contributed to the final version of the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks Lewis Roberts and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Comparison of CLCA with other HCC cohorts.
a, Comparison of clinical information between CLCA and PCAWG-HCC. DP, double positive of HBV and HCV; DN, double negative of HBV and HCV. b, Sequencing depth of 494 tumours and their matched normal controls in CLCA. c, Relationships among driver genes using the DISCOVER mutual exclusivity test. d-e, Venn plot showing the comparison of potential driver genes identified in the TCGA-HCC, PCAWG-HCC, and our CLCA cohort. *Potential non-true drivers curated by PCAWG-HCC. f-g, Comparison of frequency of potential drivers between CLCA and PCAWG-HCC (f) and TCGA-HCC (g), respectively. Two-sided Fisher’s exact test, multiple hypothesis test performed with the Benjamini–Hochberg method. A threshold of Q < 0.1 was used for significance and denoted in blue. h, Two-sided Spearman correlation between the ratio of clonal drivers and tumour purity across all CLCA samples. The grey shaded area represents the 95% confidence interval. i, The dN/dS ratios for clonal and subclonal SNVs in 23 cancer coding drivers across our CLCA cohort. n denotes the total number of mutations for each category collected from 494 individual tumours. Centre points denote dN/dS values for missense, nonsense, splice site, and all mutations. Error bars denote the 95% confidence intervals. Red dashed line denotes dN/dS value of 1. j, Workflow for mutational signature analysis in CLCA.
Extended Data Fig. 2 Profiles of all mutational signatures in CLCA.
Mutational profiles of all signatures. SBS (single base substitution), DBS (doublet base substitution), and ID (small insertion and deletion). Magnified versions of signatures SBS_H1, DBS_H1 and ID_H1 are shown to illustrate the classification of each mutation subtype in each plot. The cosine similarity between each signature and its matched COSMICv3.2 signature is indicated. Novel signatures are labelled in red.
Extended Data Fig. 3 Analysis of mutational signatures.
a, Signature profiles of SBS_H8, DBS_H2, and ID_H3 extracted by both mSigHdp and SigProfiler. b, Comparison of the pentanucleotide context of SBS_H8, SBS_H2, and AA-exposed cell lines. The red square highlights the pentanucleotide context of T > A mutations enriched in SBS_H8 compared to SBS_H2. c, Correlation between the numbers of mutations associated with SBS_H2, DBS_H2, and ID_H3. d, Mutational profile of DBS_H1. e, Mutational profile of ID_H8 related to aflatoxin. f, Correlation between numbers of DBS_H1 mutations and age for involved patients. g, Correlation between numbers of ID_H8 mutations and SBS_H3 for involved patients. h, Unsupervised clustering based on the proportions of SBS, DBS, and ID mutations across tumours results in five subgroups. Selected clinical variables are also listed. The P values indicate significant nonrandom distributions for each attribute. Two-sided Fisher’s exact tests with Benjamini-Hochberg correction for multiple comparison. A threshold of Q < 0.1 was used for significance. i, Bar plots comparing selected variables that had significant differences between groups. Blue denotes mutation or yes. Grey denotes wildtype or no. Two-sided Chi-square test. j, Boxplots comparing the contributions of SBS_H8 across five subgroups. n denotes biologically independent samples. For boxplots, centre line shows median, box limits indicate upper and lower quartiles, and whiskers extend 1.5 times the interquartile range, while data beyond the end of the whiskers are outlying points that are plotted individually. Two-tailed Student’s t-test. k, OS and DFS of CLCA cases stratified into SBS_H8-high and SBS_H8-low groups by the median value. Log-rank test. For c, f, and g, ρ and P values are from a two-sided Spearman correlation test.
Extended Data Fig. 4 Mutational signature attributions.
a, mSigHdp splits COSMIC SBS5 into three components: SBS_H1, SBS_H14 and SBS_H17. They together recapitulate the pattern of SBS5. SigProfiler splits COSMIC SBS5 into four components: SBS96C, SBS96E, SBS96I and SBS96M. They together recapitulate the pattern of SBS5. b, Correlation between patient age with mutation numbers of mSigHdp extracted SBS5-split signature (SBS_H1, SBS_H14 and SBS_H17) or of SigProfiler extracted SBS5-split signature (SBS96C, SBS96E, SBS96I and SBS96M). c, mSigHdp splits COSMIC SBS40 into three components: SBS_H4, SBS_H16 and SBS_H17. They together recapitulate the pattern of SBS40. SigProfiler splits COSMIC SBS40 into three components: SBS96E, SBS96M and SBS96I. They together recapitulate the pattern of SBS40. d, Correlation between patient age with mutation numbers of mSigHdp extracted SBS40-split signature (SBS_H4, SBS_H16 and SBS_H17) or of SigProfiler extracted SBS40-split signature (SBS96E, SBS96M and SBS96I). e-f, Stacked bar plots showing the contributions of SBS mutational processes, coloured as shown in Extended Data Fig. 3h, to coding driver mutations (e) and noncoding driver mutations (f). g, Stacked bar plot shows the contribution of mutational processes to hotspot mutations (chromosome: position: the total number of patients with mutations at this particular genomic hotspot). Gene names are given with amino acid alterations for protein-coding genes. h, Enrichment of mutational signatures with clonal status. Potential aetiology and related COSMIC signatures are annotated for each signature. Two-sided Chi-square test. For b and d, ρ and P values are from a two-sided Spearman correlation test.
Extended Data Fig. 5 Survival, CNAs, HBV integrations and ecDNA.
a, Multi-variate analysis for OS and DFS. Multivariate Cox analysis was performed. Hazard ratios with a 95% confidence interval are shown for each predictor and are plotted on a natural log scale. b, Significant CNAs identified by GISTIC analysis. Red for amplification and blue for deletion. Green lines denote the threshold of Q value = 0.001. c, Hotspots of HBV integrations across CLCA. d, Top frequently amplified genes detected in ecDNA. e, Boxplots comparing the copy number of genes detected in ecDNA to others. f, Higher expression of oncogenes in ecDNA compared with these not in ecDNA. In e-f, n denotes biologically independent samples. Two-sided Wilcoxon rank-sum test. For boxplots, centre line shows median, box limits indicate upper and lower quartiles, and whiskers extend 1.5 times the interquartile range, while data beyond the end of the whiskers are outlying points that are plotted individually. g, Comparison of the frequency of cases with kataegis events (denoted in blue) between patients with or without APOBEC signatures. Two-sided Chi-square test.
Extended Data Fig. 6 Patterns of SVs and clustered mutational processes.
a, The number of SV events, focal CN segments, kataegis events, chromoplexy events, and chromothripsis events in the CLCA. b, Proportions of different categories for each type of alteration. c, The density of breakpoints genome-wide (top) and 2D density of juxtapositions (bottom) of SV, chromothripsis, and chromoplexy. d-e, Examples of chromothripsis (d) and chromoplexy (e) events involving oncogenes.
Extended Data Fig. 7 Kataegis and evolutionary history.
a, Rainfall plots of kataegis events. n denotes the total number of kataegis events detected in the tumour and marked with arrows below. b, Distribution of point mutations over different mutation periods. c, Distribution of mutations across early clonal, late clonal and subclonal stages, for drivers in CLCA. Barplots comparing the distribution of coding and noncoding mutations are shown, Two-sided Chi-square test. d, Relative ordering of CN events and driver mutations across all samples. e, Relative timing of signatures across all patients. n denotes the total number of individual tumours with the presence of the noted signature. For boxplots, centre line shows median, box limits indicate upper and lower quartiles, and whiskers extend 1.5 times the interquartile range, while data beyond the end of the whiskers are outlying points that are plotted individually. Boxplots are ordered by the median and no statistical test is used. f, Preferential ordering diagrams for patients stratified based on Cluster V, alcohol, and smoking. The relative ordering of candidate drivers was compared.
Extended Data Fig. 8 Dysregulated pathways.
Each gene box includes the frequency of patients influenced by different types of somatic alterations affecting the corresponding gene. A total of eight forms of somatic alterations are listed and colour-coded, including coding SNVs, noncoding SNVs (further divided into promoters, lncRNAs and UTRs), CNAs, ecDNA, SVs and HBV integrations. Solid rectangles enclose genes in eight major signalling pathways. Dashed rectangles enclose genes in specific signalling pathways. Interactions between genes are indicated. For each pathway, the frequencies of patients altered by coding mutations only, noncoding mutations only, and both coding and noncoding mutations are denoted, as shown in the Venn diagram.
Extended Data Fig. 9 Functional validation of PPP1R12B and KCNJ12.
a, Comparison of tumour migration, invasion, self-renewal and cell proliferation capacities of PPP1R12B disruption across cell lines. b, Edited sites in PPP1R12B by Prime Editing. c, RT-qPCR analysis of PPP1R12B mRNA expression across wild-type (WT) and point-mutated HepG2 cell lines. d-e, Comparison of the proliferation (d), migration, invasion, and self-renewal (e) capacities across cell lines of indicated genotypes. Representative images of each assay are shown for each cell line. f, Comparison of tumour migration, invasion, self-renewal and cell proliferation capacities of KCNJ12 disruption across cell lines. g, Edited sites in KCNJ12. h, RT-qPCR analysis of KCNJ12 mRNA expression across wild-type (WT) and point-mutated HepG2 cell lines. i-j, Comparison of the proliferation (i), migration, invasion, and self-renewal (j) capacities across cell lines of indicated genotypes. For all panels, each experimental condition was independently repeated for three times. Representative images of each assay are shown. Data are presented as mean ± s.e.m. In a, e, f, j, P values for the comparison between a certain group with the control group are denoted on the top of images. Two-way ANOVA test is used for proliferation analysis in (a, d, f, i). For other plots, P value was derived with two-tailed Student’s t-test.
Extended Data Fig. 10 Function validation of FGA.
a, Lollipop plot of FGA mutations in CLCA. b, Overall survival of TCGA-HCC patients (n = 364) classified by FGA expression levels, Log-rank test. c, Comparison of FGA mRNA expression between tumour and normal tissues in the TCGA-HCC cohort. For boxplots, centre line shows median, box limits indicate upper and lower quartiles, and whiskers extend 1.5 times the interquartile range, while data beyond the end of the whiskers are outlying points that are plotted individually. d, Representative FGA IHC images of paired tumour and normal tissues. Quantitative result is shown in Fig. 5d. e, Schematic of the edited site in the FGA noncoding region. f, Western blot analysis of FGA levels across wild-type and mutated HepG2 cell lines. Source gels in Supplementary Fig. 3. g-h, Comparison of the proliferation (g), migration, invasion, and self-renewal (h) capacities across cell lines of indicated genotypes. i-j, Comparison of tumour migration (i), invasion and self-renewal, and cell proliferation (j) capacities of FGA disruption across cell lines. k, Resected xenograft tumours by sh-Ctrl (n = 6) and sh-FGA cells (n = 7) in PLC/PRF/5. l, Specific phospho-antibody array analysis between PLC/PRF/5-sh-Ctrl and sh-FGA cell lines. Top significantly altered phosphorylation sites among 156 phosphoproteins are listed. m, TYK2 phosphorylation and its unphosphorylated counterpart between PLC/PRF/5- sh-Ctrl and sh-FGA cell lines determined with Cy3-labelled streptavidin via specific phospho-antibody array (n = 2 for each phosphorylated site or unphosphorylated protein). n, Western blot analysis of p-TYK2 (Y1054) and p-STAT3 (Y705) protein levels by FGA knockdown in PLC/PRF/5 and PVTT cell lines. p-, phosphorylated. Source gels in Supplementary Fig. 4. o, Representative images of cell migration assay following inhibitor treatment. p, IL6 mRNA expression of sh-Ctrl and sh-FGA cells. q, IL6 mRNA levels between PLC/PRF/5- sh-Ctrl and sh-FGA cell lines following FBS stimulation. Cells were incubated in DMEM supplemented with 10% FBS for the indicated time intervals after treated with FBS-free medium overnight. r, Two-tailed Pearson correlation analysis of FGA protein and TYK2 phosphorylation (n = 75) in an independent HCC patient cohort. The relative intensity of FGA and p-TYK2 were normalized to β-actin. Source gels in Supplementary Fig. 5. s, A proposed model illustrating the role of the FGA/TYK2/STAT3 axis during HCC tumorigenesis. Wildtype and mutated forms of FGA were shown, respectively. The diagram was created using BioRender. For all panels, n denotes biologically independent samples. Each experimental condition was independently repeated three to five times. Data are presented as mean ± s.e.m. In h and i, P value for the comparison between a certain group with the control group are denoted on the top of images. Two-tailed Student’s t-test is used in (c, h, i, p, and q). Two-way ANOVA test is used in g and j.
Supplementary information
Supplementary Information
Supplementary Figs. 1–5, Supplementary Notes 1–8 and Supplementary Discussion.
Supplementary Table 1
Clinical information and mutational landscape.
Supplementary Table 2
TPM matrix of RNA-seq cases.
Supplementary Table 3
Mutational signatures.
Supplementary Table 4
Catalogue of genomic rearrangements including CNAs, SVs, HBV integrations and clustered alterations.
Supplementary Table 5
Mutational history.
Supplementary Table 6
Signalling pathways.
Supplementary Table 7
Functional validation of candidate drivers and reagents.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Chen, L., Zhang, C., Xue, R. et al. Deep whole-genome analysis of 494 hepatocellular carcinomas. Nature 627, 586–593 (2024). https://doi.org/10.1038/s41586-024-07054-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-024-07054-3
This article is cited by
-
HCC genomic landscape in Chinese individuals
Nature Reviews Gastroenterology & Hepatology (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.