Abstract
Purpose
Interpreting free-text contrast-enhanced ultrasound (CEUS) reports can lead to diagnostic errors and prolonged patient waiting times. Generative Pre-trained Transformer (GPT)-4, a state-of-the-art natural language processing model, may improve diagnostic efficiency by generating structured medical reports from unstructured data. This experimental study investigates the impact of GPT-4-generated structured reports on doctors’ diagnostic efficiency in liver nodule CEUS examinations, comparing their performance with that of doctors using conventional free-text reports.
Methods
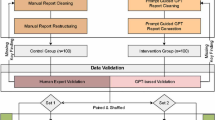
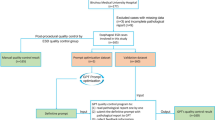
A total of 159 CEUS reports were collected and structured using GPT-4, and 30 doctors of varying experience levels participated in the study. The performance of doctors using free-text reports was compared with those using structured reports in terms of diagnostic efficiency and accuracy.
Results
The study revealed significant improvements in diagnostic efficiency (20 vs. 17 min) and accuracy (73% vs. 79%) for doctors using GPT-4-generated structured reports compared to traditional free-text reports. This trend was consistent across all experience levels. Qualitative insights from frontline ultrasound doctors provided valuable feedback on the strengths and weaknesses of GPT-4-generated structured reports.
Conclusion
GPT-4-generated structured reports show potential in enhancing diagnostic efficiency and accuracy in liver nodule CEUS examinations. Despite certain limitations, refining GPT-4 or similar natural language processing models in future iterations can yield greater benefits. Future research should explore broader clinical applications and investigate GPT models and natural language processing techniques in areas such as decision support, patient communication, and medical research, ultimately contributing to improved patient care and healthcare outcomes.
Clinical Relevance
This study suggests that GPT-4-generated structured reports enhance diagnostic efficiency and accuracy in liver nodule CEUS examinations, potentially improving patient care and outcomes by reducing diagnostic errors and patient wait times.



Similar content being viewed by others
Data Availability
The datasets generated and/or analyzed during the current study are not publicly available due to privacy and ethical considerations but are available from the corresponding author on reasonable request. All requests for data access will be reviewed by the author to ensure appropriate use. Further information on the data and how to request access can be obtained from the corresponding author.
Abbreviations
- CEUS:
-
Contrast-enhanced ultrasound
- GPT:
-
Generative pre-trained transformer
- AIGC:
-
AI-generated content
- NLP:
-
Natural language processing
References
Georgiou, A., Li, J., Thomas, J., Dahm, M. R., & Westbrook, J. I. (2019). The impact of health information technology on the management and follow-up of test results—a systematic review. Journal of the American Medical Informatics Association, 26(7), 678–688.
Awaysheh, A., Wilcke, J., Elvinger, F., Rees, L., Fan, W., & Zimmerman, K. J. (2018). A review of medical terminology standards and structured reporting. Journal of Veterinary Diagnostic Investigation, 30(1), 17–25.
Johnson, A. J., Chen, M. Y., Zapadka, M. E., Lyders, E. M., & Littenberg, B. (2010). Radiology report clarity: A cohort study of structured reporting compared with conventional dictation. Journal of the American College of Radiology, 7(7), 501–506.
Bink, A., Benner, J., Reinhardt, J., De Vere-Tyndall, A., Stieltjes, B., Hainc, N., et al. (2018). Structured reporting in neuroradiology: Intracranial tumors. Frontiers in Neurology, 9, 32.
Brown, T., Mann, B., Ryder, N., Subbiah, M., Kaplan, J. D., Dhariwal, P., et al. (2020). Language models are few-shot learners. Advances in Neural Information Processing Systems, 33, 1877–1901.
Adams, L. C., Truhn, D., Busch, F., Kader, A., Niehues, S. M., Makowski, M. R., et al. (2023). Leveraging GPT-4 for post hoc transformation of free-text radiology reports into structured reporting: A multilingual feasibility study. Radiology, 307, e230725.
Wang, P., Berzin, T. M., Brown, J. R. G., Bharadwaj, S., Becq, A., Xiao, X., et al. (2019). Real-time automatic detection system increases colonoscopic polyp and adenoma detection rates: A prospective randomised controlled study. Gut, 68(10), 1813–1819.
Névéol, A., Dalianis, H., Velupillai, S., Savova, G., & Zweigenbaum, P. (2018). Clinical Natural Language Processing in languages other than English: Opportunities and challenges. Journal of Biomedical Semantics, 9(1), 12.
Xue, K., Zhou, Y., Ma, Z., Ruan, T., Zhang, H., & He, P. (Eds.) Fine-tuning BERT for joint entity and relation extraction in Chinese medical text. In 2019 IEEE International Conference on Bioinformatics and Biomedicine (BIBM); 2019. IEEE.
Kono, Y., Lyshchik, A., Cosgrove, D., Dietrich, C. F., Jang, H.-J., Kim, T. K., et al. (2017). Contrast enhanced ultrasound (CEUS) liver imaging reporting and data system (LI-RADS®): The official version by the American College of Radiology (ACR). Ultraschall in der Medizin-European Journal of Ultrasound, 38(01), 85–86.
Radiology. ACo. CEUS LI-RADS®v2017 CORE.
Gosbee, J. (2002). Human factors engineering and patient safety. Quality & Safety in Health Care, 11(4), 352–354.
Jiang, M., Chen, Y., Liu, M., Rosenbloom, S. T., Mani, S., Denny, J. C., et al. (2011). A study of machine-learning-based approaches to extract clinical entities and their assertions from discharge summaries. Journal of the American Medical Informatics Association, 18(5), 601–606.
Radford, A., Narasimhan, K., Salimans, T., & Sutskever, I. (2018). Improving language understanding by generative pre-training.
Wang, Y., Sohn, S., Liu, S., Shen, F., Wang, L., Atkinson, E. J., et al. (2019). A clinical text classification paradigm using weak supervision and deep representation. BMC Medical Informatics and Decision Making, 19, 1–13.
Langlotz, C. P. J. R. (2009). Structured radiology reporting: Are we there yet? Radiology, 253(1), 23–5.
Ganeshan, D., Duong, P.-A.T., Probyn, L., Lenchik, L., McArthur, T. A., Retrouvey, M., et al. (2018). Structured reporting in radiology. Academic Radiology, 25(1), 66–73.
Dreyer, K. J., Kalra, M. K., Maher, M. M., Hurier, A. M., Asfaw, B. A., Schultz, T., et al. (2005). Application of recently developed computer algorithm for automatic classification of unstructured radiology reports: Validation study. Radiology, 234(2), 323–329.
Bosmans, J., Peremans, L., Menni, M., De Schepper, A., Duyck, P., & Parizel, P. J. (2012). Structured reporting: If, why, when, how—and at what expense? Results of a focus group meeting of radiology professionals from eight countries. Insights into Imaging, 3, 295–302.
Acknowledgments
We express our sincere gratitude to the 30 medical professionals from various institutions who participated in this study.
Funding
Beijing key Clinical Discipline Funding (No. 2021-135).
Author information
Authors and Affiliations
Contributions
ZW: Conceptualization, Writing—Original Draft Preparation. GR, SP, QL: Data Collection, Data Analysis. XH: Supervision, Review & Editing of the Manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no conflicts of interest to declare.
Ethical Approval
The study was approved by our institutional review board. Informed consent was waived due to the retrospective and anonymous nature of our study.
Consent to Publish
Before the submission of our manuscript, we have obtained consent to publish from all individuals who participated in our study. This includes consent for the publication of anonymized data and any other potentially identifiable information included in the manuscript. We understand that the responsibility for ensuring this consent lies with us, the authors, and we have taken all necessary steps to safeguard the privacy and rights of the participants. We have also ensured that all participants have been informed about the nature of the publication and the potential use of the data in the public domain.
Consent to Participate
All participants involved in this study were fully informed about the nature, objectives, potential benefits, and risks of their participation. Written informed consent was obtained from each participant prior to their involvement in the study. In cases where participants were under the age of 18, consent was obtained from a parent or legal guardian. We ensured that participation in this study was completely voluntary, with participants having the right to withdraw at any point without any negative consequences. The process of obtaining consent was conducted in accordance with the ethical guidelines and approvals set forth by our institutional review board.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wang, Z., Guo, R., Sun, P. et al. Enhancing Diagnostic Accuracy and Efficiency with GPT-4-Generated Structured Reports: A Comprehensive Study. J. Med. Biol. Eng. 44, 144–153 (2024). https://doi.org/10.1007/s40846-024-00849-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40846-024-00849-9