Abstract
Background
Epigenetics contributes to the pathogenesis of amyotrophic lateral sclerosis (ALS). We aimed to characterize the DNA methylation profiles associated with clinical heterogeneity in disease progression and survival among patients.
Methods
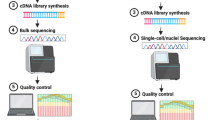
We included a cohort of 41 patients with sporadic ALS, with a median follow-up of 86.9 months, and 27 rigorously matched healthy controls. Blood-based genome-wide DNA methylation analysis was conducted.
Results
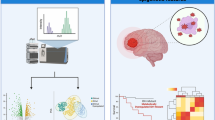
A total of 948 progression rate-associated differentially methylated positions, 298 progression rate-associated differentially methylated regions (R-DMRs), 590 survival time-associated DMPs, and 197 survival time-associated DMRs (S-DMRs) were identified, using complementary grouping strategies. Enrichment analysis of differentially methylated genes highlighted the involvement of synapses and axons in ALS progression and survival. Clinical analysis revealed a positive correlation between the average methylation levels of the R-DMR in PRDM8 and disease progression rate (r = 0.479, p = 0.002). Conversely, there was an inverse correlation between the average methylation levels of the R-DMR in ANKRD33 and disease progression rate (r = − 0.476, p = 0.002). In addition, patients with higher methylation levels within the S-DMR of ZNF696 experienced longer survival (p = 0.016), while those with elevated methylation levels in the S-DMR of RAI1 had shorter survival (p = 0.006).
Conclusion
DNA methylation holds promise as a potential biomarker for tracking disease progression and predicting survival outcome and also offers targets for precision medicine.


Similar content being viewed by others
Availability of data and materials
Data and materials are available from the corresponding author on reasonable request.
Abbreviations
- ABLIM1:
-
Actin-binding LIM protein 1
- AGRN:
-
Agrin
- ALS:
-
Amyotrophic lateral sclerosis
- sALS:
-
Sporadic amyotrophic lateral sclerosis
- ALSFRS-R:
-
Revised-Amyotrophic Lateral Sclerosis Functional Rating Scale
- ANKRD33:
-
Ankyrin repeat domain 33
- BRSK2:
-
BR serine/threonine kinase 2
- CTTN:
-
Cortactin
- C9orf72:
-
Chromosome 9 open reading frame 72
- DLG2:
-
Discs large MAGUK scaffold protein 2
- DNM1:
-
Dynamin 1
- DSS:
-
Dispersion shrinkage for sequencing data
- FUS:
-
Fused in sarcoma
- GO:
-
Gene ontology
- HC:
-
Healthy controls
- ITGB1:
-
Integrin subunit beta 1
- LYN:
-
LYN proto-oncogene
- MC-seq:
-
Methylation capture bisulfite sequencing
- PIEZO2:
-
Piezo-type mechanosensitive ion channel component 2
- PLXND1:
-
Plexin D1
- PPI:
-
Protein–protein interaction
- PRDM8:
-
PR/SET domain 8
- PTK2:
-
Protein tyrosine kinase 2
- RAI1:
-
Retinoic acid induced 1
- R-DMPs:
-
Progression rate-associated differentially methylated positions
- R-DMRs:
-
Progression rate-associated differentially methylated regions
- S-DMPs:
-
Survival time-associated differentially methylated positions
- S-DMRs:
-
Survival time-associated differentially methylated regions
- ZNF696:
-
Zinc finger protein 696
References
Feldman EL, Goutman SA, Petri S et al (2022) Amyotrophic lateral sclerosis. Lancet 400:1363
Zarei S, Carr K, Reiley L et al (2015) A comprehensive review of amyotrophic lateral sclerosis. Surg Neurol Int 6:171
Li C, Liu J, Lin J, Shang H (2022) COVID-19 and risk of neurodegenerative disorders: a Mendelian randomization study. Transl Psychiatry 12:283
Yang T, Wei Q, Li C et al (2022) Spatial-temporal pattern of propagation in amyotrophic lateral sclerosis and effect on survival: a cohort study. Eur J Neurol 29:3177–3186
Hussain N (2012) Epigenetic influences that modulate infant growth, development, and disease. Antioxid Redox Signal 17:224–236
Machnik M, Oleksiewicz U (2020) Dynamic signatures of the epigenome: friend or foe? Cells 9:653
Hillary RF, McCartney DL, Smith HM et al (2023) Blood-based epigenome-wide analyses of 19 common disease states: a longitudinal, population-based linked cohort study of 18,413 Scottish individuals. PLoS Med 20:e1004247
Hop PJ, Zwamborn RAJ, Hannon E et al (2022) Genome-wide study of DNA methylation shows alterations in metabolic, inflammatory, and cholesterol pathways in ALS. Sci Transl Med 14:0264
Ruf WP, Hannon E, Freischmidt A et al (2022) Methylome analysis of ALS patients and presymptomatic mutation carriers in blood cells. Neurobiol Aging 116:16–24
Cai Z, Jia X, Liu M, Yang X, Cui L (2022) Epigenome-wide DNA methylation study of whole blood in patients with sporadic amyotrophic lateral sclerosis. Chin Med J (Engl) 135:1466–1473
Martin LJ, Adams DA, Niedzwiecki MV, Wong M (2022) Aberrant DNA and RNA methylation occur in spinal cord and skeletal muscle of human SOD1 mouse models of ALS and in human ALS: targeting DNA methylation is therapeutic. Cells 11:3448
Appleby-Mallinder C, Schaber E, Kirby J et al (2021) TDP43 proteinopathy is associated with aberrant DNA methylation in human amyotrophic lateral sclerosis. Neuropathol Appl Neurobiol 47:61–72
Cedarbaum JM, Stambler N, Malta E et al (1999) The ALSFRS-R: a revised ALS functional rating scale that incorporates assessments of respiratory function. BDNF ALS Study Group (Phase III). J Neurol Sci 169:13–21
Wang J, C. P, M. N, inventors; Agilent SureSelectXT Methyl-Seq applications with low-input DNA and smaller capture libraries. https://www.agilent.com/cs/library/applications/5991-7838EN.pdf2017. Accessed 24 June 2023
Shu C, Zhang X, Aouizerat BE, Xu K (2020) Comparison of methylation capture sequencing and Infinium MethylationEPIC array in peripheral blood mononuclear cells. Epigenet Chromatin 13:51
Krueger F, Andrews SR (2011) Bismark: a flexible aligner and methylation caller for Bisulfite-Seq applications. Bioinformatics 27:1571–1572
Ryan DP, Ehninger D (2014) Bison: bisulfite alignment on nodes of a cluster. BMC Bioinform 15:337
Akalin A, Kormaksson M, Li S et al (2012) methylKit: a comprehensive R package for the analysis of genome-wide DNA methylation profiles. Genome Biol 13:R87
Teh AL, Pan H, Lin X et al (2016) Comparison of Methyl-capture Sequencing vs. Infinium 450K methylation array for methylome analysis in clinical samples. Epigenetics 11:36–48
Park Y, Wu H (2016) Differential methylation analysis for BS-seq data under general experimental design. Bioinformatics 32:1446–1453
Wu T, Hu E, Xu S et al (2021) clusterProfiler 4.0: a universal enrichment tool for interpreting omics data. Innovation (Camb) 2:100141
Szklarczyk D, Kirsch R, Koutrouli M et al (2023) The STRING database in 2023: protein-protein association networks and functional enrichment analyses for any sequenced genome of interest. Nucl Acids Res 51:D638–D646
McIntyre JC, Titlow WB, McClintock TS (2010) Axon growth and guidance genes identify nascent, immature, and mature olfactory sensory neurons. J Neurosci Res 88:3243–3256
Orozco D, Edbauer D (2013) FUS-mediated alternative splicing in the nervous system: consequences for ALS and FTLD. J Mol Med (Berl) 91:1343–1354
Wang Y, Wan B, Li D et al (2012) BRSK2 is regulated by ER stress in protein level and involved in ER stress-induced apoptosis. Biochem Biophys Res Commun 423:813–818
Nothling J, Abrahams N, Toikumo S et al (2021) Genome-wide differentially methylated genes associated with posttraumatic stress disorder and longitudinal change in methylation in rape survivors. Transl Psychiatry 11:594
Cypris O, Eipel M, Franzen J et al (2020) PRDM8 reveals aberrant DNA methylation in aging syndromes and is relevant for hematopoietic and neuronal differentiation. Clin Epigenet 12:125
Maierhofer A, Flunkert J, Oshima J et al (2019) Epigenetic signatures of Werner syndrome occur early in life and are distinct from normal epigenetic aging processes. Aging Cell 18:e12995
Jiang L, Penney KL, Giovannucci E, Kraft P, Wilson KM (2018) A genome-wide association study of energy intake and expenditure. PLoS ONE 13:e0201555
Ludolph A, Dupuis L, Kasarskis E, Steyn F, Ngo S, McDermott C (2023) Nutritional and metabolic factors in amyotrophic lateral sclerosis. Nat Rev Neurol 19:511
Woo SH, Lukacs V, de Nooij JC et al (2015) Piezo2 is the principal mechanotransduction channel for proprioception. Nat Neurosci 18:1756–1762
Sonkodi B (2023) Miswired proprioception in amyotrophic lateral sclerosis in relation to pain sensation (and in delayed onset muscle soreness)—is piezo2 channelopathy a principal transcription activator in proprioceptive terminals besides being the potential primary damage? Life (Basel) 13:657
Chang YT, Kowalczyk M, Fogerson PM et al (2022) Loss of Rai1 enhances hippocampal excitability and epileptogenesis in mouse models of Smith-Magenis syndrome. Proc Natl Acad Sci USA 119:e2210122119
Fogarty MJ, Noakes PG, Bellingham MC (2015) Motor cortex layer V pyramidal neurons exhibit dendritic regression, spine loss, and increased synaptic excitation in the presymptomatic hSOD1(G93A) mouse model of amyotrophic lateral sclerosis. J Neurosci 35:643–647
Fogarty MJ, Klenowski PM, Lee JD et al (2016) Cortical synaptic and dendritic spine abnormalities in a presymptomatic TDP-43 model of amyotrophic lateral sclerosis. Sci Rep 6:37968
Gulino R (2023) Synaptic dysfunction and plasticity in amyotrophic lateral sclerosis. Int J Mol Sci 24:4613
Verma M, Lizama BN, Chu CT (2022) Excitotoxicity, calcium and mitochondria: a triad in synaptic neurodegeneration. Transl Neurodegener 11:3
Fogarty MJ (2019) Amyotrophic lateral sclerosis as a synaptopathy. Neural Regen Res 14:189–192
Traynelis SF, Wollmuth LP, McBain CJ et al (2010) Glutamate receptor ion channels: structure, regulation, and function. Pharmacol Rev 62:405–496
Yang S, Park JH, Lu HC (2023) Axonal energy metabolism, and the effects in aging and neurodegenerative diseases. Mol Neurodegener 18:49
Okamoto K, Hirai S, Shoji M, Senoh Y, Yamazaki T (1990) Axonal swellings in the corticospinal tracts in amyotrophic lateral sclerosis. Acta Neuropathol 80:222–226
Chia R, Chio A, Traynor BJ (2018) Novel genes associated with amyotrophic lateral sclerosis: diagnostic and clinical implications. Lancet Neurol 17:94–102
Jones PA (2012) Functions of DNA methylation: islands, start sites, gene bodies and beyond. Nat Rev Genet 13:484–492
Zhang L, Silva TC, Young JI et al (2020) Epigenome-wide meta-analysis of DNA methylation differences in prefrontal cortex implicates the immune processes in Alzheimer’s disease. Nat Commun 11:6114
Chuang YH, Paul KC, Bronstein JM, Bordelon Y, Horvath S, Ritz B (2017) Parkinson’s disease is associated with DNA methylation levels in human blood and saliva. Genome Med 9:76
Wei QQ, Chen YP, Chen XP et al (2018) Prognostic nomogram associated with longer survival in amyotrophic lateral sclerosis patients. Aging Dis 9:965–975
Braun PR, Han S, Hing B et al (2019) Genome-wide DNA methylation comparison between live human brain and peripheral tissues within individuals. Transl Psychiatry 9:47
Acknowledgements
The authors thank the patients and their families for their participation in the study.
Funding
This study was supported by the National Natural Science Foundation of China (Grant nos. 82371430 and 82101485) and Sichuan Science and Technology Program (Grant no. 2022ZDZX0023).
Author information
Authors and Affiliations
Contributions
TMY and HFS conceived and designed the study. TMY, CYL, QQW, YFC, JXH, JYL, YX, QRJ, and SCW collected the data. TMY, CYL, and DJP contributed to the statistical analysis. TMY wrote the first draft of the manuscript. TMY, HFS, CYL, and QQW contributed to the writing of the final version of the manuscript. DJP provided critical review. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflicts of interest
The authors declare that they have no conflict of interest.
Ethics approval and consent to participate
Ethics approval for the study was approved by the institutional ethics committee of Sichuan University. Written informed consent was obtained from each participant or their primary caregiver.
Consent for publication
Not applicable.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yang, T., Li, C., Wei, Q. et al. Genome-wide DNA methylation analysis related to ALS patient progression and survival. J Neurol 271, 2672–2683 (2024). https://doi.org/10.1007/s00415-024-12222-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00415-024-12222-6